
Acta Horticulturae Sinica ›› 2023, Vol. 50 ›› Issue (3): 475-484.doi: 10.16420/j.issn.0513-353x.2021-1267
• Research Papers • Previous Articles Next Articles
NING Yuansheng, LI Huan, SONG Jianfei, YU Tingting, HAN Mengyuan, PENG Lulin, JIA Junqi, ZHANG Weiwei, YANG Hongqiang*( )
)
Received:2022-06-17
Revised:2022-11-09
Online:2023-03-25
Published:2023-04-03
Contact:
*(E-mail:hqyang@sdau.edu.cn)
CLC Number:
NING Yuansheng, LI Huan, SONG Jianfei, YU Tingting, HAN Mengyuan, PENG Lulin, JIA Junqi, ZHANG Weiwei, YANG Hongqiang. Characterization of NCL Family Genes in Malus and Their Relationship with Cellular Calcium Concentration in Root[J]. Acta Horticulturae Sinica, 2023, 50(3): 475-484.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2021-1267
| 基因名称 Gene name | 正向引物(5′-3′) Forward primer sequence | 反向引物(5′-3′) Revers primer sequence |
|---|---|---|
| MhNCL-1 | GGTGAACCTGATGCAGACAT | TCAGGCTTACTGATACGGGA |
| MhNCL-2 | GTCTAACAACTCCACTGTGC | GCAATAACTCGCTCCCATTC |
| MhNCL-3 | AGCTTCTCAACAGCTACGAG | ACCAGACCCAAGAATACTGC |
| MhNCL-4 | CAAGTTTGGTAAGATTCCGGG | CTGTTTCGTCCCATGTCCTT |
| MdActin | ATTCAAGTATGCCTGGGTGC | CAGTCAGCCTGTGATGTTCC |
Table 1 Primers of qRT-PCR
| 基因名称 Gene name | 正向引物(5′-3′) Forward primer sequence | 反向引物(5′-3′) Revers primer sequence |
|---|---|---|
| MhNCL-1 | GGTGAACCTGATGCAGACAT | TCAGGCTTACTGATACGGGA |
| MhNCL-2 | GTCTAACAACTCCACTGTGC | GCAATAACTCGCTCCCATTC |
| MhNCL-3 | AGCTTCTCAACAGCTACGAG | ACCAGACCCAAGAATACTGC |
| MhNCL-4 | CAAGTTTGGTAAGATTCCGGG | CTGTTTCGTCCCATGTCCTT |
| MdActin | ATTCAAGTATGCCTGGGTGC | CAGTCAGCCTGTGATGTTCC |
| 物种 Species | 基因名称 Gene name | 基因登录号 Gene ID | 氨基酸 aa | 等电点 pI | 分子量/kD Molecular weight | 染色体 Chromosome | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|
| 苹果 | MdNCL-1 | MD03G1238600 | 586 | 5.39 | 63.75 | Chr3 | 细胞膜 Cell membrane |
| Malus × domestica | MdNCL-2 | MD03G1238700 | 600 | 5.30 | 66.03 | Chr3 | 细胞膜 Cell membrane |
| MdNCL-3 | MD11G1258400 | 584 | 5.01 | 63.26 | Chr11 | 细胞膜 Cell membrane | |
| MdNCL-4 | MD17G1233800 | 585 | 5.34 | 64.56 | Chr17 | 细胞膜 Cell membrane | |
| 拟南芥 Arabidopsis thaliana | AtNCL | AT1G53210.1 | 585 | 5.15 | 63.42 | Chr1 | 细胞膜 Cell membrane |
| 葡萄 | VvNCL-1 | VIT_212s0028g02570 | 577 | 5.73 | 62.98 | Chr12 | 细胞膜 Cell membrane |
| Vitis vinifera | VvNCL-2 | VIT_219s0014g01740 | 579 | 5.21 | 63.10 | Chr19 | 细胞膜 Cell membrane |
| 草莓 | FvNCL-1 | FvH4_3g18760 | 578 | 5.75 | 63.67 | fvb3 | 细胞膜 Cell membrane |
| Fragaria vesca | FvNCL-2 | FvH4_3g18770 | 580 | 5.41 | 63.51 | fvb3 | 细胞膜 Cell membrane |
| FvNCL-3 | FvH4_6g26990 | 580 | 4.81 | 63.58 | fvb3 | 细胞膜 Cell membrane | |
| 桃 | PpNCL-1 | Prupe.3G125200 | 575 | 5.56 | 63.45 | pp03 | 细胞膜 Cell membrane |
| Prunus persica | PpNCL-2 | Prupe.4G171300 | 593 | 5.39 | 64.77 | pp04 | 细胞膜 Cell membrane |
| PpNCL-3 | Prupe.4G171400 | 579 | 5.30 | 63.26 | pp04 | 细胞膜 Cell membrane | |
| 白梨 | PbNCL-1 | Pbr038243.1 | 585 | 5.05 | 63.61 | Chr11 | 细胞膜 Cell membrane |
| Pyrus bretschneideri | PbNCL-2 | Pbr022502.1 | 585 | 5.23 | 64.46 | Chr17 | 细胞膜 Cell membrane |
| PbNCL-3 | Pbr002291.1 | 549 | 5.26 | 60.01 | scaffold1094.0 | 细胞膜 Cell membrane | |
| PbNCL-4 | Pbr002292.1 | 580 | 5.18 | 63.69 | scaffold1094.0 | 细胞膜 Cell membrane | |
| PbNCL-5 | Pbr002293.1 | 580 | 5.18 | 63.69 | scaffold1094.0 | 细胞膜 Cell membrane | |
| PbNCL-6 | Pbr002294.1 | 549 | 5.26 | 60.01 | scaffold1094.0 | 细胞膜 Cell membrane |
Table 2 The information of NCL family members from different species
| 物种 Species | 基因名称 Gene name | 基因登录号 Gene ID | 氨基酸 aa | 等电点 pI | 分子量/kD Molecular weight | 染色体 Chromosome | 亚细胞定位 Subcellular localization |
|---|---|---|---|---|---|---|---|
| 苹果 | MdNCL-1 | MD03G1238600 | 586 | 5.39 | 63.75 | Chr3 | 细胞膜 Cell membrane |
| Malus × domestica | MdNCL-2 | MD03G1238700 | 600 | 5.30 | 66.03 | Chr3 | 细胞膜 Cell membrane |
| MdNCL-3 | MD11G1258400 | 584 | 5.01 | 63.26 | Chr11 | 细胞膜 Cell membrane | |
| MdNCL-4 | MD17G1233800 | 585 | 5.34 | 64.56 | Chr17 | 细胞膜 Cell membrane | |
| 拟南芥 Arabidopsis thaliana | AtNCL | AT1G53210.1 | 585 | 5.15 | 63.42 | Chr1 | 细胞膜 Cell membrane |
| 葡萄 | VvNCL-1 | VIT_212s0028g02570 | 577 | 5.73 | 62.98 | Chr12 | 细胞膜 Cell membrane |
| Vitis vinifera | VvNCL-2 | VIT_219s0014g01740 | 579 | 5.21 | 63.10 | Chr19 | 细胞膜 Cell membrane |
| 草莓 | FvNCL-1 | FvH4_3g18760 | 578 | 5.75 | 63.67 | fvb3 | 细胞膜 Cell membrane |
| Fragaria vesca | FvNCL-2 | FvH4_3g18770 | 580 | 5.41 | 63.51 | fvb3 | 细胞膜 Cell membrane |
| FvNCL-3 | FvH4_6g26990 | 580 | 4.81 | 63.58 | fvb3 | 细胞膜 Cell membrane | |
| 桃 | PpNCL-1 | Prupe.3G125200 | 575 | 5.56 | 63.45 | pp03 | 细胞膜 Cell membrane |
| Prunus persica | PpNCL-2 | Prupe.4G171300 | 593 | 5.39 | 64.77 | pp04 | 细胞膜 Cell membrane |
| PpNCL-3 | Prupe.4G171400 | 579 | 5.30 | 63.26 | pp04 | 细胞膜 Cell membrane | |
| 白梨 | PbNCL-1 | Pbr038243.1 | 585 | 5.05 | 63.61 | Chr11 | 细胞膜 Cell membrane |
| Pyrus bretschneideri | PbNCL-2 | Pbr022502.1 | 585 | 5.23 | 64.46 | Chr17 | 细胞膜 Cell membrane |
| PbNCL-3 | Pbr002291.1 | 549 | 5.26 | 60.01 | scaffold1094.0 | 细胞膜 Cell membrane | |
| PbNCL-4 | Pbr002292.1 | 580 | 5.18 | 63.69 | scaffold1094.0 | 细胞膜 Cell membrane | |
| PbNCL-5 | Pbr002293.1 | 580 | 5.18 | 63.69 | scaffold1094.0 | 细胞膜 Cell membrane | |
| PbNCL-6 | Pbr002294.1 | 549 | 5.26 | 60.01 | scaffold1094.0 | 细胞膜 Cell membrane |
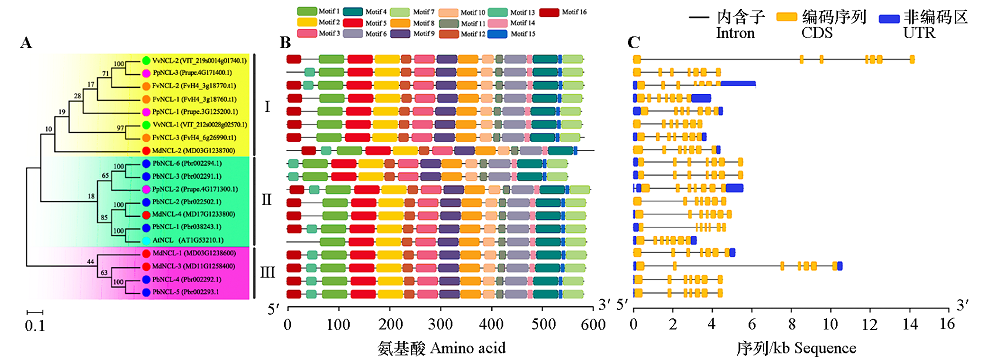
Fig. 1 Phylogenetic relationships,conserved motif and gene structure of NCL family members from different species At:Arabidopsis thaliana;Md:Malus × domestica;Fv:Fragaria vesca;Vv:Vitis vinifera;Pp:Prunus persica;Pb:Pyrus × bretschneideri.
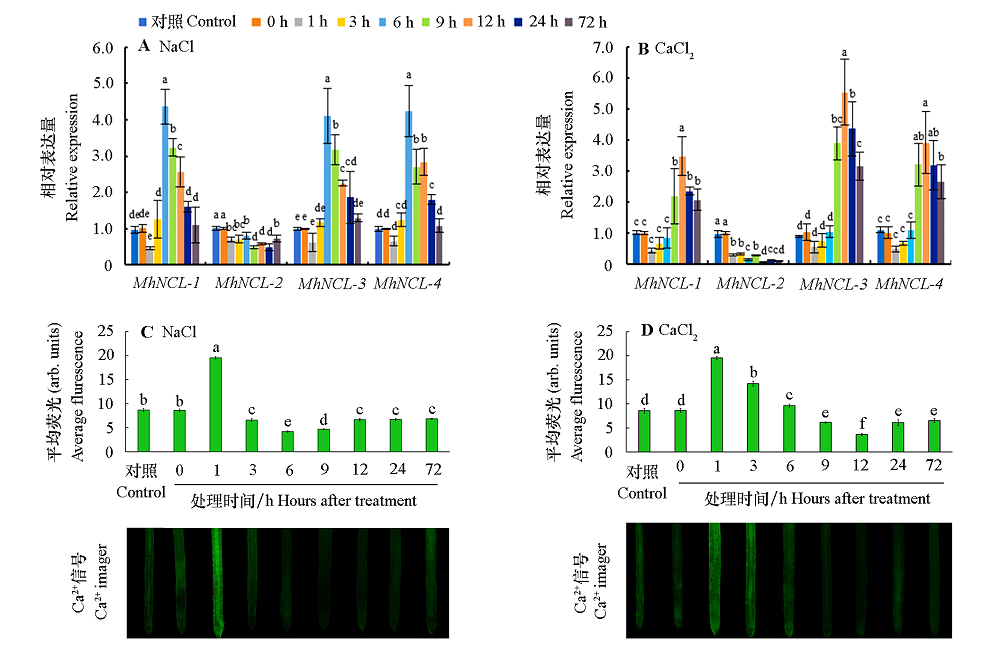
Fig. 4 Expression of MhNCL and Ca2+ concentration in root system under NaCl and CaCl2 stresses The expression level of MdNCL at 0 h was set as 1,and three replicates were set. The difference of the same gene at different treatment time was analyzed with a confidence interval of 5%(P < 0.05). The green fluorescence responds to Ca2+ concentration,the stronger the fluorescence,the higher the Ca2+ concentration,and the weaker the fluorescence,the lower the Ca2+ concentration. The same below.
| [1] |
Batistič O, Kudla J. 2012. Analysis of calcium signaling pathways in plants. Biochim Biophys Acta, 1820 (8):1283-1293.
doi: 10.1016/j.bbagen.2011.10.012 pmid: 22061997 |
| [2] | Blaustein M P, Lederer W J. 1999. Sodium/calcium exchange:its physiological implications. Physiological Reviews, 793:763-854. |
| [3] |
Chen C, Chen H, Zhang Y, Thomas H R, Frank M H, He Y, Xia R. 2020. TBtools:an integrative toolkit developed for interactive analyses of big biological data. Mol Plant, 13 (8):1194-1202.
doi: 10.1016/j.molp.2020.06.009 URL |
| [4] |
Dipolo R, Beauge L. 2006. Sodium/calcium exchanger:influence of metabolic regulation on ion carrier interactions. Physiol Rev, 86 (1):155-203.
doi: 10.1152/physrev.00018.2005 URL |
| [5] |
Do T D, Chen H, Hien V T, Hamwieh A, Yamada T, Sato T, Yan Y, Cong H, Shono M, Suenaga K, Xu D. 2016. NCL synchronously regulates Na+,K+,and Cl- in soybean and greatly increases the grain yield in saline field conditions. Sci Rep, 6:19147.
doi: 10.1038/srep19147 |
| [6] |
Dodd A N, Kudla J, Sanders D. 2010. The language of calcium signaling. Annu Rev Plant Biol, 61:593-620.
doi: 10.1146/annurev-arplant-070109-104628 pmid: 20192754 |
| [7] |
Edel K H, Marchadier E, Brownlee C, Kudla J, Hetherington A M. 2017. The evolution of calcium-based signalling in plants. Curr Biol, 27 (13):R667-R679.
doi: 10.1016/j.cub.2017.05.020 URL |
| [8] | Emery L, Whelan S, Hirschi K D, Pittman J K. 2012. Protein phylogenetic analysis of Ca2+/cation antiporters and insights into their evolution in plants. Front Plant Sci, 3:1. |
| [9] |
Hu B, Jin J P, Guo A Y, Zhang H, Luo J C, Gao G. 2015. GSDS 2.0:an upgraded gene feature visualization server. Bioinformatics, 31 (8):1296-1297.
doi: 10.1093/bioinformatics/btu817 URL |
| [10] |
Jin H Q, Xing M G, Cai C M, Li S. 2019. B-box proteins in arachis duranensis:genome-wide characterization and expression profiles analysis. Agronomy, 10 (1):1-17.
doi: 10.3390/agronomy10010001 URL |
| [11] |
Khananshvili D. 2020. Basic and editing mechanisms underlying ion transport and regulation in NCX variants. Cell Calcium, 85:102131.
doi: 10.1016/j.ceca.2019.102131 pmid: 31794905 |
| [12] |
Kumar S, Stecher G, Tamura K. 2016. MEGA7:molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol, 33 (7):1870-1874.
doi: 10.1093/molbev/msw054 URL |
| [13] | Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, Van de Peer Y, Rouzé P, Rombauts S. 2002. PlantCARE,a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res, 30 (1):325-327. |
| [14] | Letunic I, Bork P. 2018. 20 years of the SMART protein domain annotation resource. Nucleic Acids Re, 46 (D1):D493-D496. |
| [15] |
Li P H, Zhang G Y, Gonzales N, Guo Y Q, Hu H H, Park S, Zhao J. 2016. Ca2+-regulated and diurnal rhythm-regulated Na+/Ca2+ exchanger AtNCL affects flowering time and auxin signalling in Arabidopsis. Plant Cell Environ, 39 (2):377-392.
doi: 10.1111/pce.v39.2 URL |
| [16] |
Liu Y, Wang L, Xing X, Sun L P, Pan J W, Kong X P, Zhang M Y, Li D Q. 2013. ZmLEA3,a multifunctional group 3 LEA protein from maize(Zea mays L.),is involved in biotic and abiotic stresses. Plant and Cell Physiology, 54 (6):944-959.
doi: 10.1093/pcp/pct047 URL |
| [17] |
Mao K, Yang J, Wang M, Liu H Y, Guo X, Zhao S, Dong Q L, Ma F W. 2021. Genome-wide analysis of the apple CaCA superfamily reveals that MdCAX proteins are involved in the abiotic stress response as calcium transporters. BMC Plant Biology, 21 (1):81.
doi: 10.1186/s12870-021-02866-1 pmid: 33557757 |
| [18] |
Marchler-Bauer A, Bo Y, Han L Y, He J, Lanczycki C J, Lu S N, Chitsaz F, Derbyshire M K, Geer R C, Gonzales N R, Gwadz M, Hurwitz D I, Lu F, Marchler G H, Song J S, Thanki N, Wang Z X, Yamashita R A, Zhang D C, Zheng C J, Geer L Y, Bryant S H. 2017. CDD/SPARCLE:functional classification of proteins via subfamily domain architectures. Nucleic Acids Res, 45 (D1):D200-D203
doi: 10.1093/nar/gkw1129 URL |
| [19] |
Meng D, Li Y Y, Bai Y, Li M J, Cheng L L. 2016. Genome-wide identification and characterization of wrky transcriptional factor family in apple and analysis of their responses to waterlogging and drought stress. Plant Physiology and Biochemistry, 103:71-83.
doi: 10.1016/j.plaphy.2016.02.006 pmid: 26970718 |
| [20] |
Pittman J K, Hirschi K D. 2016. CAX-ing a wide net:cation/H+ transporters in metal remediation and abiotic stress signalling. Plant Biology, 18 (5):741-749.
doi: 10.1111/plb.12460 pmid: 27061644 |
| [21] | Qiu L, Hu S, Wang Y, Qu H. 2021. Accumulation of abnormal amyloplasts in pulp cells induces bitter pit in Malus domestica. Front Plant Sci, 23 (12):738726. |
| [22] |
Singh A K, Kumar R, Tripathi A K, Gupta B K, Pareek A, Singla-Pareek S L. 2015. Genome-wide investigation and expression analysis of sodium/calcium exchanger gene family in rice and Arabidopsis. Rice, 8 (1):54.
doi: 10.1186/s12284-015-0054-5 pmid: 26134707 |
| [23] | Song J F, Yang F, Xun M, Xu L X, Yang H Q. 2020. Genome-wide identification and characterization of vacuolar processing enzyme gene family and diverse expression under stress in apple(Malus × domestica). Front Plant Sci, 26;11:626. |
| [24] |
Spalding E P, Harper J F. 2011. The ins and outs of cellular Ca2+transport. Curr Opin Plant Biol, 14 (6):715-720.
doi: 10.1016/j.pbi.2011.08.001 pmid: 21865080 |
| [25] |
Su Y L, He H H, Wang P, Ma Z H, Mao J, Chen B H. 2021. Genome-wide characterization and expression analyses of the auxin/indole-3-acetic acid Aux/IAA gene family in apple Malus × domestica. Gene, 768:145302.
doi: 10.1016/j.gene.2020.145302 URL |
| [26] |
Tian Y, Dong Q L, Ji Z R, Chi F M, Cong P H, Zhou Z S. 2015. Genome-wide identification and analysis of the MADS-box gene family in apple. Gene, 555:277-290.
doi: 10.1016/j.gene.2014.11.018 pmid: 25447908 |
| [27] |
Wang P, Li Z W, Wei J S, Zhao Z L, Sun D Y, Cui S. 2012. A Na+/Ca2+exchanger-like protein AtNCL involved in salt stress in Arabidopsis. Journal of Biological Chemistry, 287 (53):44062-44070.
doi: 10.1074/jbc.M112.351643 URL |
| [28] |
Wu Q, Huang L P, Su N N, Shabala L, Wang H Y, Huang X, Wen R Y, Yu M, Cui J, Shabala S. 2020. Calcium-dependent hydrogen peroxide mediates hydrogen-rich water-reduced cadmium uptake in plant roots. Plant Physiol, 183 (3):1331-1344.
doi: 10.1104/pp.20.00377 pmid: 32366640 |
| [1] | LIU Youxian, LI Guofang, TAN Ming, YANG Zhichang, ZHOU Shiwei, HUO Wenjing, ZHANG He, SUN Jianshe, and SHAO Jianzhu. Expression Analysis of MdTOPP13/28 During Axillary Bud Outgrowth in Malus [J]. Acta Horticulturae Sinica, 2023, 49(4): 697-712. |
| [2] | YU Tingting, LI Huan, NING Yuansheng, SONG Jianfei, PENG Lulin, JIA Junqi, ZHANG Weiwei, YANG Hongqiang. Genome-wide Identification of GRAS Gene Family in Apple and Expression Analysis of Its Response to Auxin [J]. Acta Horticulturae Sinica, 2023, 50(2): 397-409. |
| [3] | HAN Xiaolei, ZHANG Caixia, LIU Kai, YANG An, YAN Jiadi, LI Wuxing, KANG Liqun, and CONG Peihua. A New Mid-ripening Apple Cultivar‘Zhongping Youlei’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 1-2. |
| [4] | SUN Simiao, WANG Kun, GAO Yuan, WANG Dajiang, and LI Lianwen. A New Ornamental Crabapple Cultivar‘Zichen’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 267-268. |
| [5] | HAN Xiaolei, ZHANG Caixia, LIU Kai, YAN Jiadi, LI Wuxing, KANG Liqun, and CONG Peihua. A New Mid-ripening Apple Cultivar‘Pingyou 2’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 1-2. |
| [6] | WANG Qiang, CONG Peihua, and LIU Xiaofeng. A New Late Ripening Apple Cultivar‘Huayou Tianwa’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 3-4. |
| [7] | WANG Qiang, CONG Peihua, and LIU Xiaofeng. A New Mid-ripening Apple Cultivar‘Huayou Baomi’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 5-6. |
| [8] | YANG Ling, CONG Peihua, WANG Qiang, LI Wuxing, and KANG Liqun. A New Mid-ripening Apple Cultivar‘Huafeng’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 7-8. |
| [9] | LIU Chuanhe, HE Han, SHAO Xuehua, LAI Duo, KUANG Shizi, XIAO Weiqiang, LIU Yan. A New Pineapple Cultivar‘Yuetong’ [J]. Acta Horticulturae Sinica, 2022, 49(9): 2053-2054. |
| [10] | GAO Yanlong, WU Yuxia, ZHANG Zhongxing, WANG Shuangcheng, ZHANG Rui, ZHANG De, WANG Yanxiu. Bioinformatics Analysis of Apple ELO Gene Family and Its Expression Analysis Under Low Temperature Stress [J]. Acta Horticulturae Sinica, 2022, 49(8): 1621-1636. |
| [11] | LIU Chaoyang, LIAO Zhichan, LU Xinxin, HE Yehua. Identification of CslD Gene Family in Pineapple and Functional Analysis of AcoCslD2a [J]. Acta Horticulturae Sinica, 2022, 49(8): 1650-1662. |
| [12] | ZHENG Xiaodong, XI Xiangli, LI Yuqi, SUN Zhijuan, MA Changqing, HAN Mingsan, LI Shaoxuan, TIAN Yike, WANG Caihong. Effects and Regulating Mechanism of Exogenous Brassinosteroids on the Growth of Malus hupehensis Under Saline-alkali Stress [J]. Acta Horticulturae Sinica, 2022, 49(7): 1401-1414. |
| [13] | XIA Yan, HUANG Song, WU Xueli, LIU Yiqi, WANG Miaomiao, SONG Chunhui, BAI Tuanhui, SONG Shangwei, PANG Hongguang, JIAO Jian, ZHENG Xianbo. Identification and Analysis of Apple Viruses Diseases Based on Virome Sequencing Technology [J]. Acta Horticulturae Sinica, 2022, 49(7): 1415-1428. |
| [14] | LIU Zhaoxia, ZHANG Xin, WANG Lu, MA Yuting, CHEN Qian, ZHU Zhanling, GE Shunfeng, JIANG Yuanmao. Effects of Fertilizer Hole Application Sites on Fine Root Growth,15N Absorption and Utilization,Yield and Quality of Apple Trees [J]. Acta Horticulturae Sinica, 2022, 49(7): 1545-1556. |
| [15] | MA Weifeng, LI Yanmei, MA Zonghuan, CHEN Baihong, MAO Juan. Identification of Apple POD Gene Family and Functional Analysis of MdPOD15 Gene [J]. Acta Horticulturae Sinica, 2022, 49(6): 1181-1199. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd