
Acta Horticulturae Sinica ›› 2022, Vol. 49 ›› Issue (11): 2325-233.doi: 10.16420/j.issn.0513-353x.2021-0940
• Research Papars • Previous Articles Next Articles
DAI Wenshan, WU Yue, WANG Min( )
)
Received:2022-03-30
Revised:2022-08-25
Online:2022-11-25
Published:2022-11-25
Contact:
WANG Min
E-mail:wangmin0624@foxmail.com
CLC Number:
DAI Wenshan, WU Yue, WANG Min. Preliminary Study on Mechanisms of Resistance to Citrus Canker of Kumquat FcRGA1[J]. Acta Horticulturae Sinica, 2022, 49(11): 2325-233.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2021-0940
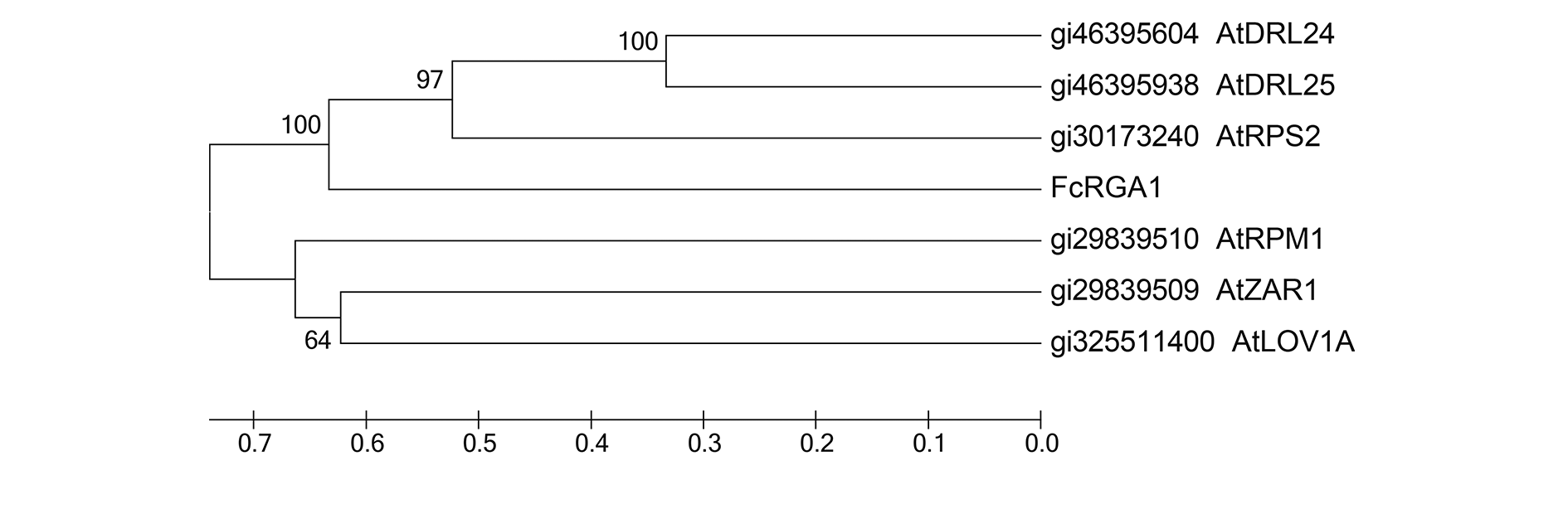
Fig. 2 Amino acid sequences phylogenetic tree of FcRGA1 and Arabidopsis(At)R protein containing NB-ARC domain The abscissa is a distance scale,representing the unit length of difference between sequences,corresponding a distance of 10 changes per 100 amino acid positions,and the numbers at node represent the distance of the kinship.
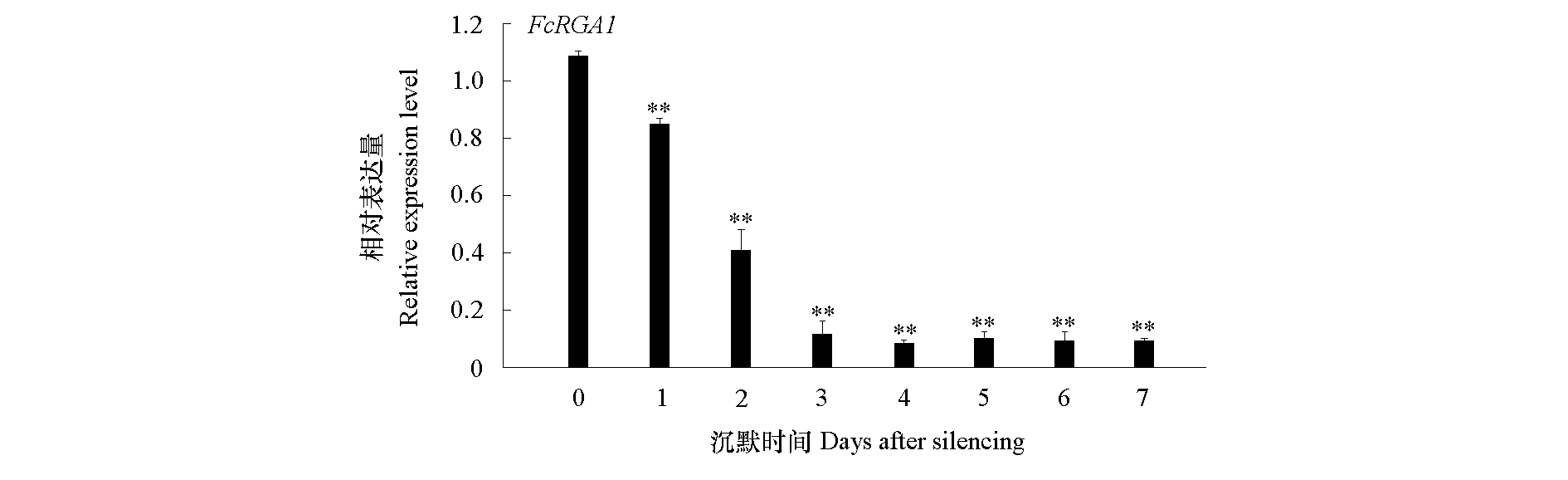
Fig. 4 Expression analysis of the FcRGA1 in the FcRGA1-silencing kumquat leaves The gene expression at each time of the control was taken as 1 for normalization. Asterisks indicate significant difference levels,**P < 0.01.

Fig. 5 The phenotype(a),fluorescence observation(b)and bacterial count(c)of control and FcRGA1-silencing kumquat leaves inoculated with Xcc C:Control;R:FcRGA1-VIGS;dpi:Days post inoculation. Asterisks indicate significant difference between FcRGA1-VIGS and control at the same time point. *P < 0.05,**P < 0.01.
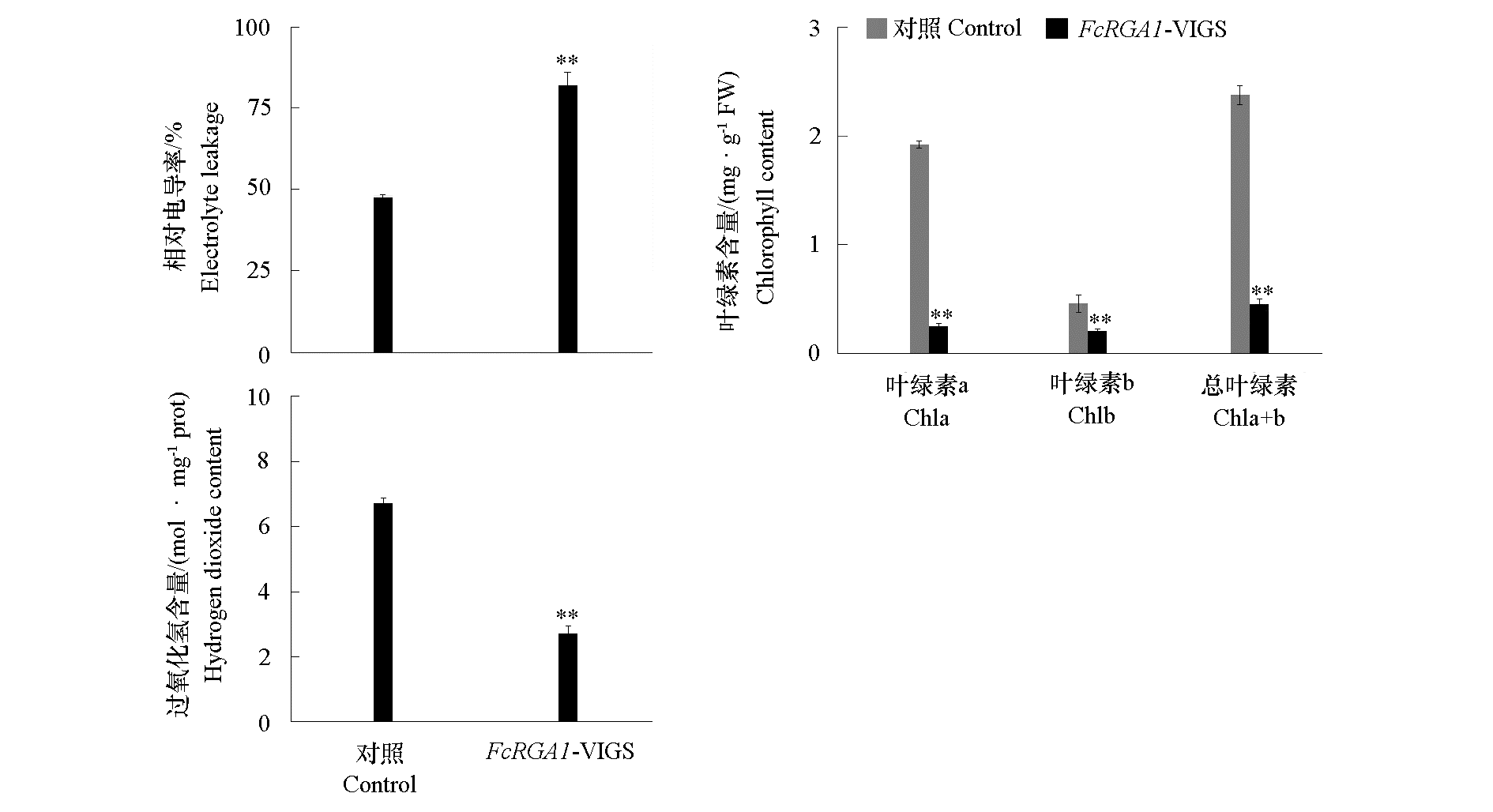
Fig. 6 Electrolyte leakage,chlorophyll content and H2O2 content of control and FcRGA1-silencing kumquat leaves after inoculated with Xcc Asterisks indicate significant difference levels,**P < 0.01.
| [1] |
Bergelson J, Kreitman M, Stahl E A, Tian D. 2001. Evolutionary dynamics of plant R-genes. Science, 292 (5525):2281-2285.
pmid: 11423651 |
| [2] | Bhure S S, Bramhankar S B, Thakur K D, Wasnik D G, Pawar R D, Labhasetwar A A, Kakad S A, Ravali T, Sarode C A. 2019. Physiological and biochemical characterization of Xanthomonas axonopodis pv. citri:a gram negative bacterium causing citrus canker. International Journal of Chemical Studies, 7 (1):1941-1944. |
| [3] |
Chenna R, Sugawara H, Koike T, Lopez R, Gibson T J, Higgins D G, Thompson J D. 2003. Multiple sequence alignment with the Clustal series of programs. Nucleic Acids Research, 31 (13):3497-3500.
doi: 10.1093/nar/gkg500 pmid: 12824352 |
| [4] |
Cui Y N, Jiang J B, Yang H H, Zhao T T, Xu X Y, Li J F. 2018. Virus-induced gene silencing (VIGS) of the NBS-LRR gene SLNLC1compromises Sm-mediated disease resistance to Stemphylium lycopersici in tomato. Biochemical and Biophysical Research Communications, 503 (3):1524-1529.
doi: 10.1016/j.bbrc.2018.07.074 URL |
| [5] |
Das A K. 2003. Citrus canker-a review. Journal of Applied Horticulture, 5 (1):52-60.
doi: 10.37855/jah.2003.v05i01.15 URL |
| [6] |
Di Lorenzo F, Silipo A, Andersen Gersby L B, Palmigiano A, Lanzetta R, Garozzo D, Boyer C, Pruvost O, Newman M, Molinaro A. 2017. Xanthomonas citri pv. citri pathotypes:LPS structure and function as microbe‐associated molecular patterns. ChemBioChem, 18 (8):772-781.
doi: 10.1002/cbic.201600671 URL |
| [7] |
Dodds P N, Rathjen J P. 2010. Plant immunity:towards an integrated view of plant-pathogen interactions. Nature Reviews Genetics, 11 (8):539-548.
doi: 10.1038/nrg2812 URL |
| [8] |
Duvaud S, Gabella C, Lisacek F, Stockinger H, Ioannidis V, Durinx C. 2021. Expasy,the Swiss Bioinformatics Resource Portal,as designed by its users. Nucleic Acids Research, 49:W216-W227.
doi: 10.1093/nar/gkab225 URL |
| [9] |
Fu X Z, Gong X Q, Zhang Y X, Wang Y, Liu J H. 2012. Different transcriptional response to Xanthomonas citri subsp. citri between kumquat and sweet orange with contrasting canker tolerance. PLoS ONE, 7:e41790.
doi: 10.1371/journal.pone.0041790 URL |
| [10] |
He H G, Zhu S Y, Zhao R H, Jiang Z N, Ji Y Y, Ji J, Qiu D, Li H J, Bie T D. 2018. Pm21,encoding a typical CC-NBS-LRR protein,confers broad-spectrum resistance to wheat powdery mildew disease. Molecular Plant, 11:879-882.
doi: 10.1016/j.molp.2018.03.004 URL |
| [11] |
Hu B, Jin J P, Guo A Y, Zhang H, Luo J C, Gao G. 2015. GSDS 2.0:an upgraded gene feature visualization server. Bioinformatics, 31(8):1296-1297.
doi: 10.1093/bioinformatics/btu817 URL |
| [12] |
Huang H, Ullah F, Zhou D X, Yi M, Zhao Y. 2019. Mechanisms of ROS regulation of plant development and stress responses. Frontiers in Plant Science, 10:800.
doi: 10.3389/fpls.2019.00800 pmid: 31293607 |
| [13] |
Khan W A, Atiq M, Sahi S T, Khan A A, Ali S, Younas M, Ali Y, Bashir M R, Sajid M. 2018. Induction of resistance against citrus canker through chemicals and plant activators. Advances in Zoology and Botany, 6 (1):26-30.
doi: 10.13189/azb.2018.060103 URL |
| [14] |
Krasileva K V, Dahlbeck D, Staskawicz B J. 2010. Activation of an Arabidopsis resistance protein is specified by the in planta association of its leucine-rich repeat domain with the cognate oomycete effector. The Plant Cell, 22 (7):2444-2458.
doi: 10.1105/tpc.110.075358 pmid: 20601497 |
| [15] |
Kun X, Zhu H F, Zhu X, Liu Z H, Wang Y, Pu W J, Guan P Y, Hu J F. 2021. Overexpression of PsoRPM3,an NBS-LRR gene isolated from myrobalan plum,confers resistance to Meloidogyne incognita in tobacco. Plant Molecular Biology, 107 (3):129-146.
doi: 10.1007/s11103-021-01185-1 URL |
| [16] |
Letunic I, Khedkar S, Bork P. 2021. SMART:recent updates,new developments and status in 2020. Nucleic Acids Research, 49 (D1):D458-D460.
doi: 10.1093/nar/gkaa937 pmid: 33104802 |
| [17] |
Li N Y, Zhou L, Zhang D D, Klosterman S J, Li T G, Gui Y J, Kong Z Q, Ma X F, Short D P G, Zhang W Q, Li J J, Subbarao K V, Chen J Y, Dai X F. 2018. Heterologous expression of the cotton NBS-LRR gene GbaNA1 enhances verticillium wilt resistance in Arabidopsis. Frontiers in Plant Science, 9:119.
doi: 10.3389/fpls.2018.00119 URL |
| [18] | Li Qiang, Qi Jingjing, Dou Wanfu, Qin Xiujuan, He Yongrui, Chen Shanchun. 2020. Overexpression of CsNBS-LRR in citrus confers bacterial canker resistance by regulating SA signaling pathway. Acta Horticulturae Sinica, 47 (5):817-826. (in Chinese) |
| 李强, 祁静静, 窦万福, 秦秀娟, 何永睿, 陈善春. 2020. 柑橘超量表达CsNBS-LRR通过SA信号途径增强对溃疡病抗性. 园艺学报, 47 (5):817-826. | |
| [19] |
Liu J H, Inoue H, Moriguchi T. 2008. Salt stress-mediated changes in free polyamine titers and expression of genes responsible for polyamine biosynthesis of apple in vitro shoots. Environmental And Experimental Botany, 62 (1):28-35.
doi: 10.1016/j.envexpbot.2007.07.002 URL |
| [20] |
Moffett P, Farnham G, Peart J, Baulcombe D C. 2002. Interaction between domains of a plant NBS-LRR protein in disease resistance-related cell death. The EMBO Journal, 21 (17):4511-4519.
doi: 10.1093/emboj/cdf453 URL |
| [21] |
Rairdan G J, Moffett P. 2006. Distinct domains in the ARC region of the potato resistance protein Rx mediate LRR binding and inhibition of activation. The Plant Cell, 18 (8):2082-2093.
doi: 10.1105/tpc.106.042747 URL |
| [22] |
Sekhwal M K, Li P C, Lam I, Wang X, Cloutier S, You F M. 2015. Disease resistance gene analogs(RGAs)in plants. International Journal of Molecular Sciences, 16 (8):19248-19290.
doi: 10.3390/ijms160819248 pmid: 26287177 |
| [23] |
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S. 2011. MEGA5:molecular evolutionary genetics analysis using maximum likelihood,evolutionary distance,and maximum parsimony methods. Molecular Biology and Evolution, 28 (10):2731-2739.
doi: 10.1093/molbev/msr121 URL |
| [24] |
van der Biezen E A, Jones J D G. 1998. Plant disease-resistance proteins and the gene-for-gene concept. Trends in Biochemical Sciences, 23 (12):454-456.
doi: 10.1016/s0968-0004(98)01311-5 pmid: 9868361 |
| [25] |
van Ooijen G, Mayr G, Kasiem M M A, Albrecht M, Cornelissen B J C, Takken F L W. 2008. Structure-function analysis of the NB-ARC domain of plant disease resistance proteins. Journal of Experimental Botany, 59 (6):1383-1397.
doi: 10.1093/jxb/ern045 pmid: 18390848 |
| [26] |
Wang C L, Zhang X P, Fan Y L, Gao Y, Zhu Q L, Zheng C K, Qin T F, Li Y Q, Che J Y, Zhang M W, Yang B, Liu Y G, Zhao K J. 2015. XA 23 is an executor R protein and confers broad-spectrum disease resistance in rice. Molecular Plant, 8 (2):290-302.
doi: 10.1016/j.molp.2014.10.010 URL |
| [27] |
Wang F S, Wang M, Liu X N, Xu Y Y, Zhu S P, Shen W X, Zhao X C. 2017. Identification of putative genes involved in limonoids biosynthesis in citrus by comparative transcriptomic analysis. Frontiers in Plant Science, 8:782.
doi: 10.3389/fpls.2017.00782 pmid: 28553308 |
| [28] |
Wang X, Chen Q, Huang J, Meng X, Cui N, Yu Y, Fan H. 2021. Nucleotide-binding leucine-rich repeat genes CsRSF1 and CsRSF2are positive modulators in the Cucumis sativus defense response to Sphaerotheca fuliginea. International Journal of Molecular Sciences, 22 (8):3986.
doi: 10.3390/ijms22083986 URL |
| [29] | Wu C H, Krasileva K V, Banfield M J, Terauchi R, Kamoun S. 2015. The“sensor domains”of plant NLR proteins:more than decoys? Frontiers in Plant Science, 6:134. |
| [30] |
Xiao X, Li X, Chen C, Guo W. 2020. DR5 is a suitable system for studying the auxin response in the Poncirus trifoliata-Xanthomonas axonopodis pv. citri interaction. Horticultural Plant Journal, 6 (5):277-283.
doi: 10.1016/j.hpj.2020.07.001 URL |
| [31] |
Xu Q, Wen X P, Deng X X. 2005. Isolation of TIR and nonTIR NBS-LRR resistance gene analogues and identification of molecular markers linked to a powdery mildew resistance locus in chestnut rose(Rosa roxburghii Tratt). Theoretical and Applied Genetics, 111 (5):819-830.
doi: 10.1007/s00122-005-0002-7 URL |
| [32] |
Xu Y J, Liu F, Zhu S W, Li X Y. 2018. The maize NBS-LRR gene ZmNBS25 enhances disease resistance in rice and Arabidopsis. Frontiers in Plant Science, 9:1033.
doi: 10.3389/fpls.2018.01033 URL |
| [33] |
Zhang C, Chen H, Cai T C, Deng Y, Zhuang R R, Zhang N, Zeng Y H, Zheng Y X, Tang R H, Pan R L, Zhuang W J. 2017. Overexpression of a novel peanut NBS-LRR gene AhRRS5 enhances disease resistance to Ralstonia solanacearum in tobacco. Plant Biotechnology Journal, 15 (1):39-55.
doi: 10.1111/pbi.12589 URL |
| [34] | Zhang Jie, Dong Shameng, Wang Wei, Zhao Jianhua, Chen Xuewei, Guo Huishan, He Guangcun, He Zuhua, Kang Zhensheng, Li Yi, Peng Youliang,Wang Guoliang,Zhou Xueping,Wang Yuanchao,Zhou Jianmin. 2019. Plant immunity research and green control of disease and insect resistance:progress,opportunities and challenges. Scientia Sinica Vitae, 49 (11):1479-1507. (in Chinese) |
| 张杰, 董莎萌, 王伟, 赵建华, 陈学伟, 郭惠珊, 何光存, 何祖华, 康振生, 李毅, 彭友良, 王国梁, 周雪平, 王源超, 周俭民. 2019. 植物免疫研究与抗病虫绿色防控:进展、机遇与挑战. 中国科学:生命科学, 49 (11):1479-1507. |
| [1] | LIU Binghao, DENG Guangzhou, TANG Zhipeng, QIN Rongyao, WEI Riji, XIA Lianghan, and QIN Qiang. A New Kumquat Cultivar‘Fuyuan Jingan’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 71-72. |
| [2] | QIU Keli, WANG Yumin, HE Jinling, YU Hong, PAN Haifa, SHENG Yu, XIE Qingmei, CHEN Hongli, ZHOU Hui, ZHANG Jinyun. Identification of Peach Laccase Family Genes and Function Analysis of PpLAC21 [J]. Acta Horticulturae Sinica, 2022, 49(6): 1351-1362. |
| [3] | XIAO Xiao, DING Liang, CONG Yu, XU Xuling, QIAO Huan, HE Silong, XU Wenjian, XU Tianshun. Isolation and Identification of a Lytic Bacteriophage Infecting Xanthomonas axonopodis pv. citri [J]. Acta Horticulturae Sinica, 2021, 48(12): 2349-2359. |
| [4] | HE Xiaowen,HE Ping,CHANG Yuansheng,WANG Sen,WANG Haibo*,and LI Linguang*. Advances in the Research on Resistance Mechanism of Apple to Ring Rot Disease [J]. ACTA HORTICULTURAE SINICA, 2020, 47(8): 1585-1594. |
| [5] | LI Qiang,QI Jingjing,DOU Wanfu,QIN Xiujuan,HE Yongrui*,and CHEN Shanchun*. Overexpression of CsNBS-LRR in Citrus Confers Bacterial Canker Resistance by Regulating SA Signaling Pathway [J]. ACTA HORTICULTURAE SINICA, 2020, 47(5): 817-826. |
| [6] | LIU Mengyu,LIU Xiaofeng,JIANG Dong,ZHU Shiping,SHEN Wanxia,YU Xin,XUE Yang,and ZHAO Xiaochun*. Identification of Genes Related to Oil Gland Development in Kumquat by Using BSA-Seq [J]. ACTA HORTICULTURAE SINICA, 2019, 46(5): 841-854. |
| [7] | WANG Lijuan,LONG Junhong,XIE Zhu,WU Liu,PENG Aihong,HE Yongrui,LONG Qin,CHEN Shanchun,and ZOU Xiuping*. Cloning and Expression Characteristics of CsWRKY22 Promoters in Response to Xanthomonas citri subsp. citri [J]. ACTA HORTICULTURAE SINICA, 2019, 46(4): 677-690. |
| [8] | SHAO Xiuhong1,2,DOU Tongxin2,LIN Xuexi1,2,SHENG Ou2,DENG Guiming2,BI Fangcheng2,HU Chunhua2,LI Chunyu2,YI Ganjun2,and YANG Qiaosong2,*. Research Status and Prospects on Resistance Mechanisms to Fusarium Wilt of Banana [J]. ACTA HORTICULTURAE SINICA, 2018, 45(9): 1661-1674. |
| [9] | QI Dongxia1,*,ZHANG Ying1,*,ZHAO Zhen1,ZHANG Weiwei1,CHEN Yuhui1,LIAN Yong1,TIAN Shibing2,and LIU Fuzhong1,**. Optimization of Virus-induced Gene Silencing System and Function Identification of SmIAA19 Gene in Eggplant [J]. ACTA HORTICULTURAE SINICA, 2018, 45(4): 691-701. |
| [10] | LIAO Jingjing,NIU Congcong,XIE Qunjie,XING Qiaojuan,and QI Hongyan*. The Recent Advances of Transient Expression System in Horticultural Plants [J]. ACTA HORTICULTURAE SINICA, 2017, 44(9): 1787-1795. |
| [11] | CHENG Chunzhen1,CHEN Dengyun2,LIU Weihua1,LIN Yuling1,ZHUO Chunxuan3,LIU Shengcai1,CHEN Yukun1,ZHANG Zihao1,and LAI Zhongxiong1,*. A New and Early-ripening Kumquat Cultivar‘Jinqiuzao’ [J]. ACTA HORTICULTURAE SINICA, 2017, 44(4): 803-804. |
| [12] | YANG Sha*,ZHANG Bin*,HAN Yao,YANG Xia,LI Mingyang,and GUO Yulong**. Cloning of Petunia PDS Gene and Its Application in shRNA-mediated Gene Silencing [J]. ACTA HORTICULTURAE SINICA, 2017, 44(2): 315-322. |
| [13] | JIA Ruirui,HU Anhua,CHEN Shanchun,ZOU Xiuping,PENG Aihong,XU Lanzhen,LEI Tiangang,YAO Lixiao,BAI Xiaojing,HE Yongrui*,and LI Qiang*. Cloning and Expression Analysis of CsAP-09:a Transcription Factor Related to Citrus Canker Disease [J]. ACTA HORTICULTURAE SINICA, 2017, 44(10): 1881-1893. |
| [14] | YAO Ting-shan1,2,ZHOU Yan1,and ZHOU Chang-yong1,2,*. Advances in Copper Resistant Mechanisms and Control Methods of Citrus Canker [J]. ACTA HORTICULTURAE SINICA, 2016, 43(9): 1711-1719. |
| [15] | YAO Ting-shan,ZHOU Yan,and ZHOU Chang-yong. Research Development of the Differentiation and Control of Citrus Bacterial Canker Disease [J]. ACTA HORTICULTURAE SINICA, 2015, 42(9): 1699-1706. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd