
Acta Horticulturae Sinica ›› 2021, Vol. 48 ›› Issue (12): 2360-2374.doi: 10.16420/j.issn.0513-353x.2020-0911
Previous Articles Next Articles
HOU Qiandong1, SHEN Tianjiao1, YU Huanhuan1, QIU Zhilang1, WEN Zhuang1, ZHANG Huimin1,2, WU Yawei3, WEN Xiaopeng1,*( )
)
Received:2021-01-06
Revised:2021-05-06
Published:2022-01-04
Contact:
WEN Xiaopeng
E-mail:xpwensc@hotmail.com
CLC Number:
HOU Qiandong, SHEN Tianjiao, YU Huanhuan, QIU Zhilang, WEN Zhuang, ZHANG Huimin, WU Yawei, WEN Xiaopeng. Genome-wide Identification and Expression Analysis of Prunus avium Gretchen Hagen 3(GH3)Gene Family[J]. Acta Horticulturae Sinica, 2021, 48(12): 2360-2374.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2020-0911
| 基因 Gene | 正向引物(5′-3′) Forward primer | 反向引物(5′-3′) Reverse primer | 扩增产物/bp PCR product | 退火温度/℃ Tm |
|---|---|---|---|---|
| PavGH3.1 | CAGTTGCAGAAATGCTGTGTCC | TCATGTGGCTTGCTTAGTGGGT | 276 | 61 |
| PavGH3.2 | GGTCCTCAGCAGCCGAATCAC | CCACCCTTCAGAAGCGCCATA | 265 | 64 |
| PavGH3.3 | ACTTCTTTGGTGCCCCTTGCTTC | GTGGTCTGTGCGCTATGGTGTGT | 179 | 65 |
| PavGH3.4 | CTCTTACACCCTCATTCCCTCCAT | CGACTCTCAGCACATCTCCAACT | 224 | 59 |
| PavGH3.5 | CATAGAGGAGACGACCAGGAACA | GGAACTCAGAGATGGGATGAGAGCT | 239 | 62 |
| PavGH3.6 | CTCCCATCTTGTCAGCTCACCCA | GCTCACGAACAGAAAGTACAGGCCC | 197 | 62 |
| PavGH3.7 | CTGTTGAGGTTCTTGGTGGGC | AGTTTTCTCGGGGGGTTGCA | 263 | 60 |
| PavGH3.8 | CTCCCTTGCTTTCTCTGCCTAAT | GACCACAGCCTCAATGTACTTG | 125 | 60 |
| PavEF1-α2 | ATCCAGAGTAGCAGAACCAATCAC | GTTAGGCATCCAGTCCCAGAAT | 127 | 58 |
| PavRSP3 | TCAAGGTCAGGTAAGGGGGTC | GTGAGGTGATTGTTAGTGGAAAGC | 223 | 56.5 |
Table 1 Quantitative real-time PCR primer of Prunus avium GH3 gene family
| 基因 Gene | 正向引物(5′-3′) Forward primer | 反向引物(5′-3′) Reverse primer | 扩增产物/bp PCR product | 退火温度/℃ Tm |
|---|---|---|---|---|
| PavGH3.1 | CAGTTGCAGAAATGCTGTGTCC | TCATGTGGCTTGCTTAGTGGGT | 276 | 61 |
| PavGH3.2 | GGTCCTCAGCAGCCGAATCAC | CCACCCTTCAGAAGCGCCATA | 265 | 64 |
| PavGH3.3 | ACTTCTTTGGTGCCCCTTGCTTC | GTGGTCTGTGCGCTATGGTGTGT | 179 | 65 |
| PavGH3.4 | CTCTTACACCCTCATTCCCTCCAT | CGACTCTCAGCACATCTCCAACT | 224 | 59 |
| PavGH3.5 | CATAGAGGAGACGACCAGGAACA | GGAACTCAGAGATGGGATGAGAGCT | 239 | 62 |
| PavGH3.6 | CTCCCATCTTGTCAGCTCACCCA | GCTCACGAACAGAAAGTACAGGCCC | 197 | 62 |
| PavGH3.7 | CTGTTGAGGTTCTTGGTGGGC | AGTTTTCTCGGGGGGTTGCA | 263 | 60 |
| PavGH3.8 | CTCCCTTGCTTTCTCTGCCTAAT | GACCACAGCCTCAATGTACTTG | 125 | 60 |
| PavEF1-α2 | ATCCAGAGTAGCAGAACCAATCAC | GTTAGGCATCCAGTCCCAGAAT | 127 | 58 |
| PavRSP3 | TCAAGGTCAGGTAAGGGGGTC | GTGAGGTGATTGTTAGTGGAAAGC | 223 | 56.5 |
| 基因 Gene | 基因ID Gene ID | 染色体 chromosome | CDS/bp | 氨基酸 长度/aa Length | 蛋白 分子量 Molecular weight | 理论 等电点 Theoretical pI | 亲水性 GRAVY | 亚细胞 定位预测 Subcellular localization |
|---|---|---|---|---|---|---|---|---|
| PavGH3.1 | Pav_sc0001540.1_g090.1.mk | PAV_r1.0chr1 | 1 851 | 616 | 70 875.15 | 5.68 | -0.295 | 细胞质,细胞核 Cytoplasm,nucleus |
| PavGH3.2 | Pav_sc0000254.1_g140.1.mk | PAV_r1.0chr2 | 1 683 | 560 | 62 622.79 | 5.80 | -0.136 | 叶绿体 Chloroplast |
| PavGH3.3 | Pav_sc0002360.1_g300.1.mk | PAV_r1.0chr3 | 1 794 | 597 | 67 453.09 | 6.45 | -0.212 | 叶绿体 Chloroplast |
| PavGH3.4 | Pav_sc0000269.1_g440.1.mk | PAV_r1.0chr4 | 1 845 | 614 | 69 182.12 | 5.91 | -0.272 | 叶绿体 Chloroplast |
| PavGH3.5 | Pav_sc0000049.1_g070.1.mk | PAV_r1.0chr6 | 1 800 | 599 | 67 477.05 | 5.81 | -0.265 | 叶绿体 Chloroplast |
| PavGH3.6 | Pav_sc0001422.1_g120.1.mk | PAV_r1.0chr8 | 1 803 | 600 | 67 636.32 | 5.56 | -0.258 | 叶绿体 Chloroplast |
| PavGH3.7 | Pav_sc0003065.1_g090.1.mk | PAV_r1.0chr8 | 1 725 | 574 | 64 563.26 | 5.92 | -0.096 | 叶绿体 Chloroplast |
| PavGH3.8 | Pav_sc0000848.1_g820.1.mk | PAV_r1.0chr8 | 1 806 | 601 | 68 250.59 | 5.87 | -0.196 | 叶绿体 Chloroplast |
Table 2 Member information of PavGH3 gene family in Prunus avium
| 基因 Gene | 基因ID Gene ID | 染色体 chromosome | CDS/bp | 氨基酸 长度/aa Length | 蛋白 分子量 Molecular weight | 理论 等电点 Theoretical pI | 亲水性 GRAVY | 亚细胞 定位预测 Subcellular localization |
|---|---|---|---|---|---|---|---|---|
| PavGH3.1 | Pav_sc0001540.1_g090.1.mk | PAV_r1.0chr1 | 1 851 | 616 | 70 875.15 | 5.68 | -0.295 | 细胞质,细胞核 Cytoplasm,nucleus |
| PavGH3.2 | Pav_sc0000254.1_g140.1.mk | PAV_r1.0chr2 | 1 683 | 560 | 62 622.79 | 5.80 | -0.136 | 叶绿体 Chloroplast |
| PavGH3.3 | Pav_sc0002360.1_g300.1.mk | PAV_r1.0chr3 | 1 794 | 597 | 67 453.09 | 6.45 | -0.212 | 叶绿体 Chloroplast |
| PavGH3.4 | Pav_sc0000269.1_g440.1.mk | PAV_r1.0chr4 | 1 845 | 614 | 69 182.12 | 5.91 | -0.272 | 叶绿体 Chloroplast |
| PavGH3.5 | Pav_sc0000049.1_g070.1.mk | PAV_r1.0chr6 | 1 800 | 599 | 67 477.05 | 5.81 | -0.265 | 叶绿体 Chloroplast |
| PavGH3.6 | Pav_sc0001422.1_g120.1.mk | PAV_r1.0chr8 | 1 803 | 600 | 67 636.32 | 5.56 | -0.258 | 叶绿体 Chloroplast |
| PavGH3.7 | Pav_sc0003065.1_g090.1.mk | PAV_r1.0chr8 | 1 725 | 574 | 64 563.26 | 5.92 | -0.096 | 叶绿体 Chloroplast |
| PavGH3.8 | Pav_sc0000848.1_g820.1.mk | PAV_r1.0chr8 | 1 806 | 601 | 68 250.59 | 5.87 | -0.196 | 叶绿体 Chloroplast |
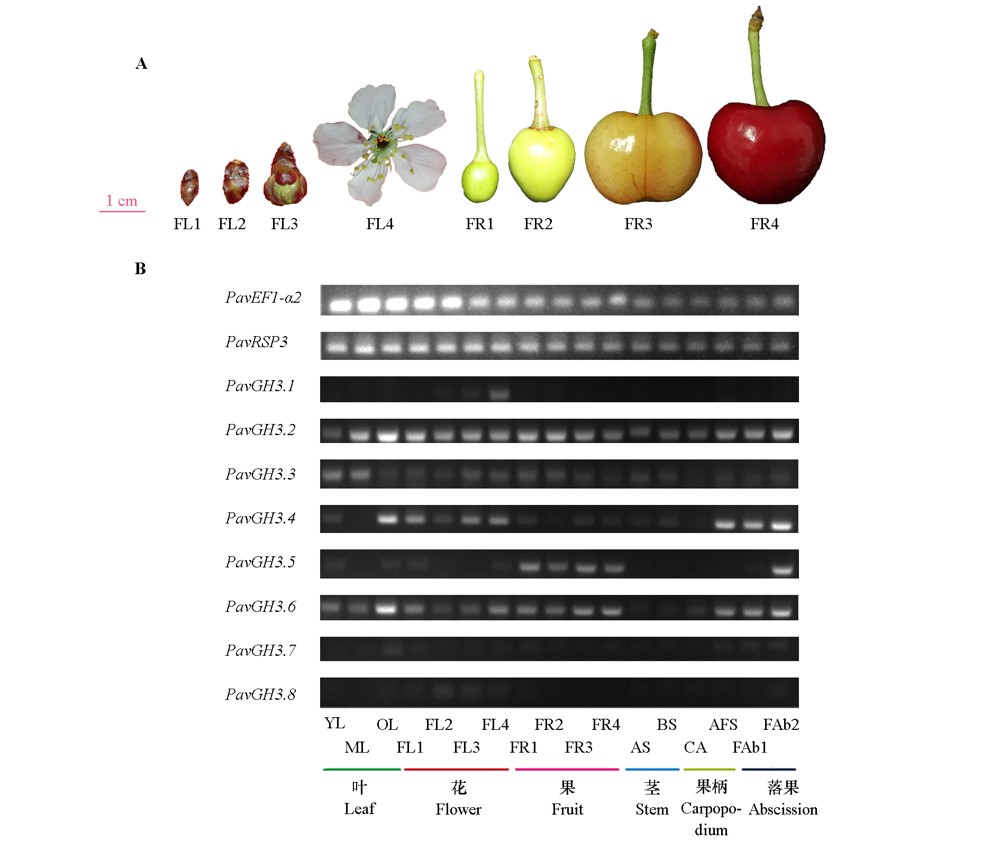
Fig. 5 PavGH3 semi-quantitative in different tissue of sweet cherries A:Flowers and fruit;B:Semi-quantitative in sweet cherries. YL:Young leaf;ML:Mature leaf;OL:Old leaf;FL1:Dormant flower bud;FL2:Flower bud;FL3:Flower bud before flowering;FL4:Blooming flowers;FR1:Small fruit;FR2:Middle fruit;FR3:Fruit of red-fleshed;FR4:Ripe fruit;AS:Annual stem;BS:Biennial stem;CA:Carpopodium;AFS:Abscising fruit carpopodium;FAb1:Fruit abscising of small fruit;FAb2:Fruit abscising of middle fruit.
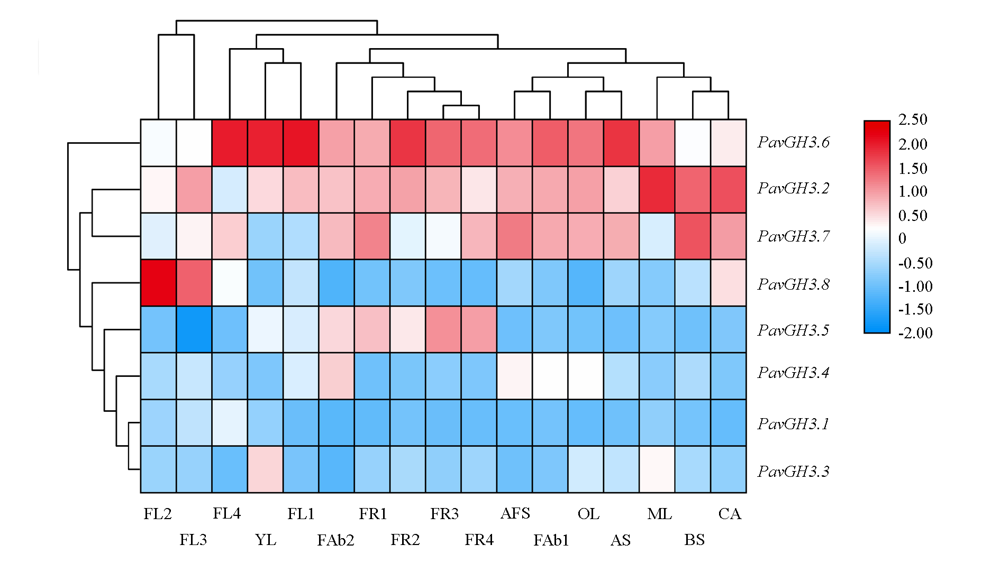
Fig. 6 PavGH3 gene expression heat map of sweet cherries YL:Young leaf;ML:Mature;OL:Old leaf;FL1:Dormant flower bud;FL2:Flower bud;FL3:Flower bud before flowering;FL4:Blooming flowers;FR1:Small fruit;FR2:Middle fruit;FR3:Fruit of red-fleshed;FR4:Ripe fruit;AS:Annual stem;BS:Biennial stem;CA:Carpopodium;AFS:Abscising fruit carpopodium;FAb1:Fruit abscising of small fruit;FAb2:Fruit abscising of middle fruit.
| [1] |
Abel S, Theologis A. 1996. Early genes and auxin action. Plant Physiology, 111 (1):9-17.
pmid: 8685277 |
| [2] | Bailey T L, Boden M, Buske F A, Frith M, Grant C E, Clementi L, Ren J, Li W W, Noble W S. 2009. MEME SUITE:tools for motif discovery and searching. Nucleic Acids Research, 37 (Web Server issue):W202-208. |
| [3] |
Berens M L, Berry H M, Mine A, Argueso C T, Tsuda K. 2017. Evolution of hormone signaling networks in plant defense. Annual Review of Plant Biology, 55:401-425.
doi: 10.1146/arplant.2004.55.issue-1 URL |
| [4] |
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden T L. 2009. BLAST+:architecture and applications. BMC Bioinformatics, 10:421.
doi: 10.1186/1471-2105-10-421 URL |
| [5] |
Casanova-Sáez R, Voß U. 2019. Auxin metabolism controls developmental decisions in land plants. Trends in Plant Science, 24 (8):741-754.
doi: S1360-1385(19)30124-4 pmid: 31230894 |
| [6] | Chao J T, Kong Y Z, Wang Q, Sun Y H, Gong D P, Lv J, Liu G S. 2015. MapGene2Chrom,a tool to draw gene physical map based on Perl and SVG languages. Yi Chuan, 37 (1):91-97. |
| [7] |
Chapman E J, Estelle M. 2009. Mechanism of auxin-regulated gene expression in plants. Annual Review of Genetics, 43:265-285.
doi: 10.1146/genet.2009.43.issue-1 URL |
| [8] |
Chen C, Chen H, Zhang Y, Thomas H R, Frank M H, He Y, Xia R. 2020. TBtools:an integrative toolkit developed for interactive analyses of big biological data. Molecular Plant, 13 (8):1194-1202.
doi: 10.1016/j.molp.2020.06.009 URL |
| [9] | Chen C Y, Ho S S, Kuo T Y, Hsieh H L, Cheng Y S. 2017. Structural basis of jasmonate-amido synthetase FIN219 in complex with glutathione S-transferase FIP 1 during the JA signal regulation. Proc Natl Acad Sci U S A, 114 (10):E1815-E1824. |
| [10] |
Chou K C, Shen H B. 2008. Cell-PLoc:a package of Web servers for predicting subcellular localization of proteins in various organisms. Nature Protocols, 3 (2):153-162.
doi: 10.1038/nprot.2007.494 URL |
| [11] |
Crooks G E, Hon G, Chandonia J M, Brenner S E. 2004. WebLogo:a sequence logo generator. Genome Research, 14 (6):1188-90.
pmid: 15173120 |
| [12] |
Denisov Y, Glick S, Zviran T, Ish-Shalom M, Levin A, Faigenboim A, Cohen Y, Irihimovitch V. 2017. Distinct organ-specific and temporal expression profiles of auxin-related genes during mango fruitlet drop. Plant Physiology and Biochemistry, 115:439-448.
doi: S0981-9428(17)30143-2 pmid: 28456120 |
| [13] | Du Yi. 2016. The Preliminary study on genes expression and and endogenous hormones quantitation analysis related to IAA and ABA during fruitlet abscission in litchi[M. D. Dissertation]. Guangzhou: South China Agricultural University. (in Chinese) |
| 杜艺. 2016. 荔枝落果过程中IAA和ABA相关基因表达及其定量检测初探[硕士论文]. 广州: 华南农业大学. | |
| [14] |
Feng L, Li G R, He Z B, Han W Y, Sun J X, Huang F L, Di J J, Chen Y S. 2019. The ARF,GH3,and Aux/IAA gene families in castor bean(Ricinus communis L.):genome-wide identification and expression profiles in high-stalk and dwarf strains. Industrial Crops and Products, 141:111804.
doi: 10.1016/j.indcrop.2019.111804 URL |
| [15] |
Feng S, Yue R, Tao S, Yang Y, Zhang L, Xu M, Wang H, Shen C. 2015. Genome-wide identification, expression analysis of auxin-responsive GH 3 family genes in maize(Zea mays L.)under abiotic stresses. Journal of Integrative Plant Biology, 57 (9):783-95.
doi: 10.1111/jipb.12327 URL |
| [16] | Guo H, Zhang Y, Wang Z, Lin L, Cui M, Long Y, Xing Z. 2019. Genome-wide identification of WRKY transcription factors in the Asteranae. Plants (Basel), 8 (10):393. |
| [17] |
Gutierrez L, Mongelard G, Floková K, Pacurar D I, Novák O, Staswick P, Kowalczyk M, Pacurar M, Demailly H, Geiss G, Bellini C. 2012. Auxin controls Arabidopsis adventitious root initiation by regulating jasmonic acid homeostasis. Plant Cell, 24 (6):2515-2527.
doi: 10.1105/tpc.112.099119 URL |
| [18] |
Hagen G, Guilfoyle T. 2002. Auxin-responsive gene expression:genes,promoters and regulatory factors. Plant Molecular Biology, 49 (3-4):373-385.
doi: 10.1023/A:1015207114117 URL |
| [19] |
Hagen G, Kleinschmidt A, Guilfoyle T. 1984. Auxin-regulated gene expression in intact soybean hypocotyl and excised hypocotyl sections. Planta, 162 (2):147-153.
doi: 10.1007/BF00410211 pmid: 24254049 |
| [20] | Jain M, Kaur N, Tyagi A K, Khurana J P. 2006. The auxin-responsive GH 3 gene family in rice (Oryza sativa). Functional & Integrative Genomics, 6 (1):36-46. |
| [21] | Khan S, Stone J M. 2007. Arabidopsis thaliana GH3.9 in auxin and jasmonate cross talk. Plant Signaling & Behavior, 2 (6):483-485. |
| [22] |
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones S J, Marra M A. 2009. Circos:an information aesthetic for comparative genomics. Genome Research, 19 (9):1639-1645.
doi: 10.1101/gr.092759.109 pmid: 19541911 |
| [23] |
Kuang J F, Wu J Y, Zhong H Y, Li C Q, Chen J Y, Lu W J, Li J G. 2012. Carbohydrate stress affecting fruitlet abscission and expression of genes related to auxin signal transduction pathway in litchi. International Journal of Molecular Sciences, 13 (12):16084-16103.
doi: 10.3390/ijms131216084 URL |
| [24] |
Kućko A, Wilmowicz E, Ostrowski M. 2019. Spatio-temporal IAA gradient is determined by interactions with ET and governs flower abscission. Journal of Plant Physiology, 236:51-60.
doi: 10.1016/j.jplph.2019.02.014 URL |
| [25] |
Kumar R, Agarwal P, Tyagi A K, Sharma A K. 2012. Genome-wide investigation and expression analysis suggest diverse roles of auxin-responsive GH 3 genes during development and response to different stimuli in tomato (Solanum lycopersicum). Mol Genet Genomics, 287 (3):221-235.
doi: 10.1007/s00438-011-0672-6 URL |
| [26] |
Kühn N, Serrano A, Abello C, Arce A, Espinoza C, Gouthu S, Deluc L, Arce-Johnson P. 2016. Regulation of polar auxin transport in grapevine fruitlets(Vitis vinifera L.)and the proposed role of auxin homeostasis during fruit abscission. BMC Plant Biology, 16 (1):234.
doi: 10.1186/s12870-016-0914-1 URL |
| [27] | Lescot M, Déhais P, Thijs G, Marchal K, Moreau Y, van de Peer Y, Rouzé P, Rombauts S. 2002. PlantCARE,a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Research, 30 (1):325-327. |
| [28] |
Ludwig-Müller J, Jülke S, Bierfreund N M, Decker E L, Reski R. 2009. Moss(Physcomitrella patens)GH 3 proteins act in auxin homeostasis. New Phytologist, 181 (2):323-338.
doi: 10.1111/j.1469-8137.2008.02677.x pmid: 19032442 |
| [29] | Madeira F, Park Y M, Lee J, Buso N, Gur T, Madhusoodanan N, Basutkar P, Tivey A R N, Potter S C, Finn R D, Lopez R. 2019. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Research, 47 (W1):W636-W641. |
| [30] |
Mockaitis K, Estelle M. 2008. Auxin receptors and plant development:a new signaling paradigm. Annual Review of Cell and Developmental Biology, 24:55-80.
doi: 10.1146/cellbio.2008.24.issue-1 URL |
| [31] |
Okrent R A, Wildermuth M C. 2011. Evolutionary history of the GH 3 family of acyl adenylases in rosids. Plant Mol Biol, 76 (6):489-505.
doi: 10.1007/s11103-011-9776-y URL |
| [32] |
Ostrowski M, Jakubowska A. 2013. GH 3 expression and IAA-amide synthetase activity in pea(Pisum sativum L.)seedlings are regulated by light,plant hormones and auxinic herbicides. Journal of Plant Physiology, 170 (4):361-368.
doi: 10.1016/j.jplph.2012.10.016 pmid: 23332498 |
| [33] |
Paponov I A, Teale W, Lang D, Paponov M, Reski R, Rensing S A, Palme K. 2009. The evolution of nuclear auxin signalling. BMC Evolutionary Biology, 9:126.
doi: 10.1186/1471-2148-9-126 pmid: 19493348 |
| [34] |
Pinto R T, Freitas N C, Máximo W P F, Cardoso T B, Prudente D O, Paiva L V. 2019. Genome-wide analysis,transcription factor network approach and gene expression profile of GH3 genes over early somatic embryogenesis in Coffea spp. BMC Genomics, 20 (1):812.
doi: 10.1186/s12864-019-6176-1 URL |
| [35] | Prakash A, Jeffryes M, Bateman A, Finn R D. 2017. The HMMER Web server for protein sequence similarity search. Current Protocols in Bioinformatics, 60:3.15.1-3.15.23. |
| [36] | Qiu Zhilang, He Meiqian, Wen Zhuang, Yang Kun, Hong Yi, Wen Xiaopeng. 2020. Selection and validation of reference genes in sweet cherry flower bud at different development stages. Seed, 39 (2):37-43. (in Chinese) |
| 仇志浪, 何美乾, 文壮, 杨鵾, 洪怡, 文晓鹏. 2020. 甜樱桃花芽不同发育时期内参基因的筛选与验证. 种子, 39 (2):37-43. | |
| [37] | Robert X, Gouet P. 2014. Deciphering key features in protein structures with the new ENDscript server. Nucleic Acids Research,42 (Web Server issue):W320-W324. |
| [38] |
Schmittgen T D, Livak K J. 2008. Analyzing real-time PCR data by the comparative CT method. Nature Protocols, 3 (6):1101-1108.
pmid: 18546601 |
| [39] |
Sherp A M, Lee S G, Schraft E, Jez J M. 2018. Modification of auxinic phenoxyalkanoic acid herbicides by the acyl acid amido synthetase GH3.15 from Arabidopsis. Journal of Biological Chemistry, 293 (46):17731-17738.
doi: 10.1074/jbc.RA118.004975 URL |
| [40] |
Staswick P E, Serban B, Rowe M, Tiryaki I, Maldonado M T, Maldonado M C, Suza W. 2005. Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. Plant Cell, 17 (2):616-627.
pmid: 15659623 |
| [41] |
Staswick P E, Tiryaki I, Rowe M L. 2002. Jasmonate response locus JAR1 and several related Arabidopsis genes encode enzymes of the firefly luciferase superfamily that show activity on jasmonic,salicylic,and indole-3-acetic acids in an assay for adenylation. Plant Cell, 14 (6):1405-1415.
pmid: 12084835 |
| [42] |
Sun R, Wang S, Ma D, Li Y, Liu C. 2019. Genome-wide analysis of cotton auxin early response gene families and their roles in somatic embryogenesis. Genes (Basel), 10 (10):730.
doi: 10.3390/genes10100730 URL |
| [43] |
Terol J, Domingo C, Talón M. 2006. The GH 3 family in plants:genome wide analysis in rice and evolutionary history based on EST analysis. Gene, 371 (2):279-290.
doi: 10.1016/j.gene.2005.12.014 URL |
| [44] |
Wang C, Liu Y, Li S S, Han G Z. 2015. Insights into the origin and evolution of the plant hormone signaling machinery. Plant Physiology, 167 (3):872-886.
doi: 10.1104/pp.114.247403 URL |
| [45] |
Wei L, Yang B, Jian H, Zhang A, Liu R, Zhu Y, Ma J, Shi X, Wang R, Li J, Xu X. 2019. Genome-wide identification and characterization of Gretchen Hagen3(GH3)family genes in Brassica napus. Genome, 62 (9):597-608..
doi: 10.1139/gen-2018-0161 URL |
| [46] |
Westfall C S, Sherp A M, Zubieta C, Alvarez S, Schraft E, Marcellin R, Ramirez L, Jez JM. 2016. Arabidopsis thaliana GH3.5 acyl acid amido synthetase mediates metabolic crosstalk in auxin and salicylic acid homeostasis. Proc Natl Acad Sci USA, 113(48):13917-13922.
doi: 10.1073/pnas.1612635113 URL |
| [47] |
Westfall C S, Zubieta C, Herrmann J, Kapp U, Nanao M H, Jez JM. 2012. Structural basis for prereceptor modulation of plant hormones by GH3 proteins. Science, 336 (6089):1708-1711.
doi: 10.1126/science.1221863 pmid: 22628555 |
| [48] |
Wilkins M R, Gasteiger E, Bairoch A, Sanchez J C, Williams K L, Appel R D, Hochstrasser D F. 1999. Protein identification and analysis tools in the ExPASy server. Methods in Molecular Biology, 112:531-552.
pmid: 10027275 |
| [49] |
Woodward A W, Bartel B. 2005. Auxin:regulation,action,and interaction. Annals of Botany, 95 (5):707-735.
pmid: 15749753 |
| [50] |
Xie R, Pang S, Ma Y, Deng L, He S, Yi S, Lv Q, Zheng Y. 2015. The ARF,AUX/IAA and GH 3 gene families in citrus:genome-wide identification and expression analysis during fruitlet drop from abscission zone A. Molecular Genetics and Genomics, 290 (6):2089-105.
doi: 10.1007/s00438-015-1063-1 URL |
| [51] |
Yang Y, Yue R, Sun T, Zhang L, Chen W, Zeng H, Wang H, Shen C. 2015. Genome-wide identification,expression analysis of GH3 family genes in Medicago truncatula under stress-related hormones and Sinorhizobium meliloti infection. Applied Microbiology and Biotechnology, 99 (2):841-854.
doi: 10.1007/s00253-014-6311-5 URL |
| [52] |
Yu D, Qanmber G, Lu L, Wang L, Li J, Yang Z, Liu Z, Li Y, Chen Q, Mendu V, Li F, Yang Z. 2018. Genome-wide analysis of cotton GH 3 subfamily II reveals functional divergence in fiber development,hormone response and plant architecture. BMC Plant Biology, 18 (1):350.
doi: 10.1186/s12870-018-1545-5 URL |
| [53] |
Yuan H, Zhao K, Lei H, Shen X, Liu Y, Liao X, Li T. 2013. Genome-wide analysis of the GH3 family in apple (Malus × domestica). BMC Genomics, 14:297.
doi: 10.1186/1471-2164-14-297 URL |
| [54] | Zeng Wen-fang, Pan Lei, Niu Liang, Lu Zhen-hua, Cui Guo-chao, Wang Zhi-qiang. 2015. Bioinformatics analysis and expression of the nectarine indole-3-aceticacid-amido synthase(GH3)gene family during fruit development. Acta Horticulturae Sinica, 42 (5):833-842. (in Chinese) |
| 曾文芳, 潘磊, 牛良, 鲁振华, 崔国朝, 王志强. 2015. 桃GH3基因家族的生物信息学分析及其在果实发育中的表达. 园艺学报, 42 (5):833-842. | |
| [55] |
Zhang C, Zhang L, Wang D, Ma H, Liu B, Shi Z, Ma X, Chen Y, Chen Q. 2018. Evolutionary history of the glycoside hydrolase 3(GH3)family based on the sequenced genomes of 48 plants and identification of jasmonic acid-related GH3 proteins in Solanum tuberosum. International Journal of Molecular Sciences, 19 (7):1850.
doi: 10.3390/ijms19071850 URL |
| [56] | Zhang He. 2013. Mechanism of dwarfing effect of M9 used as rootstock or interstem for apple[Ph. D. Dissertation]. Beijing: China Agricultural University. (in Chinese) |
| 张鹤. 2013. 苹果砧木M9作自根砧或中间砧的致矮机理研究[博士论文]. 北京: 中国农业大学. | |
| [57] |
Zhao Y. 2018. Essential roles of local auxin biosynthesis in plant development and in adaptation to environmental changes. Annual Review of Plant Biology, 69:417-435.
doi: 10.1146/arplant.2018.69.issue-1 URL |
| [58] |
Zuo X H, Xu T, Qi M F, Lü S S, Li J H, Gao S, Li T L. 2012. Expression patterns of auxin-responsive genes during tomato flower pedicel abscission and potential effects of calcium. Australian Journal of Botany, 60 (1):68.
doi: 10.1071/BT10271 URL |
| [1] | YU Tingting, LI Huan, NING Yuansheng, SONG Jianfei, PENG Lulin, JIA Junqi, ZHANG Weiwei, and YANG Hongqiang. Genome-wide Identification of GRAS Gene Family in Apple and Expression Analysis of Its Response to Auxin [J]. Acta Horticulturae Sinica, 2023, 50(2): 397-409. |
| [2] | ZHAI Hanhan, ZHAI Yujie, TIAN Yi, ZHANG Ye, YANG Li, WEN Zhiliang, CHEN Haijiang. Genome-wide Identification of Peach SAUR Gene Family and Characterization of PpSAUR5 Gene [J]. Acta Horticulturae Sinica, 2023, 50(1): 1-14. |
| [3] | ZHAO Xueyan, WANG Qi, WANG Li, WANG Fangyuan, WANG Qing, LI Yan. Comparative Transcriptome Analysis of Differential Expression in Different Tissues of Corydalis yanhusuo [J]. Acta Horticulturae Sinica, 2023, 50(1): 177-187. |
| [4] | GAO Yanlong, WU Yuxia, ZHANG Zhongxing, WANG Shuangcheng, ZHANG Rui, ZHANG De, WANG Yanxiu. Bioinformatics Analysis of Apple ELO Gene Family and Its Expression Analysis Under Low Temperature Stress [J]. Acta Horticulturae Sinica, 2022, 49(8): 1621-1636. |
| [5] | LIU Jinming, GUO Caihua, YUAN Xing, KANG Chao, QUAN Shaowen, NIU Jianxin. Genome-wide Identification of Dof Family Genes and Expression Analysis Sepal Persistent and Abscission in Pear [J]. Acta Horticulturae Sinica, 2022, 49(8): 1637-1649. |
| [6] | QIU Ziwen, LIU Linmin, LIN Yongsheng, LIN Xiaojie, LI Yongyu, WU Shaohua, YANG Chao. Cloning and Functional Analysis of the MbEGS Gene from Melaleuca bracteata [J]. Acta Horticulturae Sinica, 2022, 49(8): 1747-1760. |
| [7] | ZHENG Lin, WANG Shuai, LIU Yunuo, DU Meixia, PENG Aihong, HE Yongrui, CHEN Shanchun, ZOU Xiuping. Gene Cloning and Expression Analysis of NAC Gene in Citrus in Response to Huanglongbing [J]. Acta Horticulturae Sinica, 2022, 49(7): 1441-1457. |
| [8] | MA Weifeng, LI Yanmei, MA Zonghuan, CHEN Baihong, MAO Juan. Identification of Apple POD Gene Family and Functional Analysis of MdPOD15 Gene [J]. Acta Horticulturae Sinica, 2022, 49(6): 1181-1199. |
| [9] | ZHANG Kai, MA Mingying, WANG Ping, LI Yi, JIN Yan, SHENG Ling, DENG Ziniu, MA Xianfeng. Identification of HSP20 Family Genes in Citrus and Their Expression in Pathogen Infection Responses Citrus Canker [J]. Acta Horticulturae Sinica, 2022, 49(6): 1213-1232. |
| [10] | LIANG Chen, SUN Ruyi, XIANG Rui, SUN Yimeng, SHI Xiaoxin, DU Guoqiang, WANG Li. Genome-wide Identification of Grape GRF Family and Expression Analysis [J]. Acta Horticulturae Sinica, 2022, 49(5): 995-1007. |
| [11] | XIAO Xuechen, LIU Mengyu, JIANG Mengqi, CHEN Yan, XUE Xiaodong, ZHOU Chengzhe, WU Xingjian, WU Junnan, GUO Yinsheng, YEH Kaiwen, LAI Zhongxiong, LIN Yuling. Whole-genome Identification and Expression Analysis of SNAT,ASMT and COMT Families of Melatonin Synthesis Pathway in Dimocarpus longan [J]. Acta Horticulturae Sinica, 2022, 49(5): 1031-1046. |
| [12] | GAO Weilin, ZHANG Liman, XUE Chaoling, ZHANG Yao, LIU Mengjun, ZHAO Jin. Expression of E-type MADS-box Genes in Flower and Fruits and Protein Interaction Analysis in Chinese Jujube [J]. Acta Horticulturae Sinica, 2022, 49(4): 739-748. |
| [13] | LIU Mengyu, JIANG Mengqi, CHEN Yan, ZHANG Shuting, XUE Xiaodong, XIAO Xuechen, LAI Zhongxiong, LIN Yuling. Genome-wide Identification and Expression Analysis of GDSL Esterase/Lipase Genes in Dimocarpus longan [J]. Acta Horticulturae Sinica, 2022, 49(3): 597-612. |
| [14] | JIANG Cuicui, FANG Zhizhen, ZHOU Danrong, LIN Yanjuan, YE Xinfu. Identification and Expression Analysis of Sugar Transporter Family Genes in‘Furongli’(Prunus salicina) [J]. Acta Horticulturae Sinica, 2022, 49(2): 252-264. |
| [15] | WANG Guangpeng, LIU Tongkun, XU Xinfeng, LI Zhubo, GAO Zhanyuan, HOU Xilin. Identification of LEA Family and Expression Analysis of Some Members Under Low-temperature Stress in Chinese Cabbage [J]. Acta Horticulturae Sinica, 2022, 49(2): 304-318. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd