
Acta Horticulturae Sinica ›› 2021, Vol. 48 ›› Issue (5): 873-882.doi: 10.16420/j.issn.0513-353x.2020-1829
• Research Papers • Previous Articles Next Articles
YANG Feng, YANG Qinsong, GAO Yuhao, MA Yunjing, XU Ying, TENG Yuanwen, BAI Songling*( )
)
Received:2020-10-10
Revised:2021-01-26
Online:2021-05-25
Published:2021-06-07
Contact:
BAI Songling
E-mail:songlingbai@zju.edu.cn
CLC Number:
YANG Feng, YANG Qinsong, GAO Yuhao, MA Yunjing, XU Ying, TENG Yuanwen, BAI Songling. Establishment of Dual-cut CRISPR/Cas9 Gene Editing System in Pear Calli[J]. Acta Horticulturae Sinica, 2021, 48(5): 873-882.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.ahs.ac.cn/EN/10.16420/j.issn.0513-353x.2020-1829

Fig. 1 Illustration of the target sites and sequences of CRISPR/Cas9 in GUS gene The blue bars represent the exons,the gray bar represents the intron,the green and orange bars represent two target sites,the numbers below the bar are position information,and the width of the bars are drawn in proportion to the sequence length. The sequences below the bars are the sequences of the target sites,which are shown in the direction of 5′-3′. The red font represents the PAM sequences,and the“+/-”in parentheses represent the DNA strand recognized by the nuclease.
| 用途 Usage | 引物名称 Primer name | 引物序列(5′-3′) Sequence |
|---|---|---|
| 靶点接头正反向引物 The sense and reverse primers for the target with the adapters | GUSAtU3dT1-F | GTCATTGAGGTCGAAGACGCCAC |
| GUSAtU3dT1-R | AAACGTGGCGTCTTCGACCTCAA | |
| GUSAtU3bT2-F | GTCACGATAGCACCCTCCCGGTG | |
| GUSAtU3bT2-R | AAACCACCGGGAGGGTGCTATCG | |
| 第1轮PCR 1st PCR | U-F | CTCCGTTTTACCTGTGGAATCG |
| gR-R | CGGAGGAAAATTCCATCCAC | |
| 第2轮PCR 2nd PCR | Pps-R | TTCAGAGGTCTCTACCGACTAGTATGGAATCGGCAGCAAAGG |
| Pgs-2 | AGCGTGGGTCTCGTCAGGGTCCATCCACTCCAAGCTC | |
| Pps-2 | TTCAGAGGTCTCTCTGACACTGGAATCGGCAGCAAAGG | |
| Pgs-L | AGCGTGGGTCTCGCTCGACGCGTATCCATCCACTCCAAGCTC | |
| 载体测序引物 The sequencing primers for the vector | SP-R | TGCAATAACTTCGTATAGGCT |
| SP-L2 | GTCGTGCTCCACATGTTGACCG | |
| 阳性转化体筛选引物 The screening primers for positive transformants | Cas9-F | CTGACGCTAACCTCGACAAG |
| Cas9-R | CCGATCTAGTAACATAGATGACACC | |
| 基因编辑鉴定引物 The primers for identifying the gene editing events | GUS-F | ATGGTAGATCTGAGGGTAAATTTCTAGT |
| GUS-R | GTTCTTGTAGCCGAAATCTGGAATG |
Table 1 All primers used in this study
| 用途 Usage | 引物名称 Primer name | 引物序列(5′-3′) Sequence |
|---|---|---|
| 靶点接头正反向引物 The sense and reverse primers for the target with the adapters | GUSAtU3dT1-F | GTCATTGAGGTCGAAGACGCCAC |
| GUSAtU3dT1-R | AAACGTGGCGTCTTCGACCTCAA | |
| GUSAtU3bT2-F | GTCACGATAGCACCCTCCCGGTG | |
| GUSAtU3bT2-R | AAACCACCGGGAGGGTGCTATCG | |
| 第1轮PCR 1st PCR | U-F | CTCCGTTTTACCTGTGGAATCG |
| gR-R | CGGAGGAAAATTCCATCCAC | |
| 第2轮PCR 2nd PCR | Pps-R | TTCAGAGGTCTCTACCGACTAGTATGGAATCGGCAGCAAAGG |
| Pgs-2 | AGCGTGGGTCTCGTCAGGGTCCATCCACTCCAAGCTC | |
| Pps-2 | TTCAGAGGTCTCTCTGACACTGGAATCGGCAGCAAAGG | |
| Pgs-L | AGCGTGGGTCTCGCTCGACGCGTATCCATCCACTCCAAGCTC | |
| 载体测序引物 The sequencing primers for the vector | SP-R | TGCAATAACTTCGTATAGGCT |
| SP-L2 | GTCGTGCTCCACATGTTGACCG | |
| 阳性转化体筛选引物 The screening primers for positive transformants | Cas9-F | CTGACGCTAACCTCGACAAG |
| Cas9-R | CCGATCTAGTAACATAGATGACACC | |
| 基因编辑鉴定引物 The primers for identifying the gene editing events | GUS-F | ATGGTAGATCTGAGGGTAAATTTCTAGT |
| GUS-R | GTTCTTGTAGCCGAAATCTGGAATG |
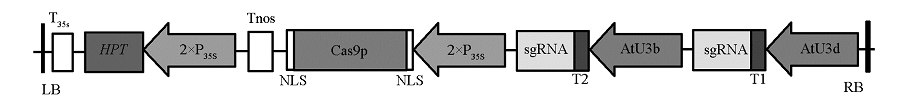
Fig. 2 Schematic diagram of GUS -pYLCRISPR/Cas9 vector LB:Left border;RB:Right border;T35s:35S terminator;Tnos:nos terminator;HPT:hygromycin B phosphotransferase;35S:Cauliflower mosaic virus 35S promoter;Cas9p:SpCas9 gene;NLS:Nuclear localization signal;AtU3d,AtU3b:U3d,U3b promoter fromArabidopsis;T1,T2:GUS Target sequence.
| 突变类型InDel types | 碱基数/bp Number of base | GUST1 sites(%) | GUST2 sites(%) | 对应株系Line# |
|---|---|---|---|---|
| 碱基插入Insertion | 1 ~ 2 | 12(30%) | 2(7.7%) | #1、#2、#3 |
| 单碱基缺失Single base deletion | 1 | 5(12.5%) | 8(30.8%) | #1、#2、#3 |
| 小片段缺失Small fragment deletion | 2 ~ 15 | 7(17.5%) | 0 | #1、#2 |
| 大片段缺失Long fragment deletion | 625 | 16(40.0%) | 16(61.5%) | #5 |
Table 2 Indel types in GUS mutant lines
| 突变类型InDel types | 碱基数/bp Number of base | GUST1 sites(%) | GUST2 sites(%) | 对应株系Line# |
|---|---|---|---|---|
| 碱基插入Insertion | 1 ~ 2 | 12(30%) | 2(7.7%) | #1、#2、#3 |
| 单碱基缺失Single base deletion | 1 | 5(12.5%) | 8(30.8%) | #1、#2、#3 |
| 小片段缺失Small fragment deletion | 2 ~ 15 | 7(17.5%) | 0 | #1、#2 |
| 大片段缺失Long fragment deletion | 625 | 16(40.0%) | 16(61.5%) | #5 |
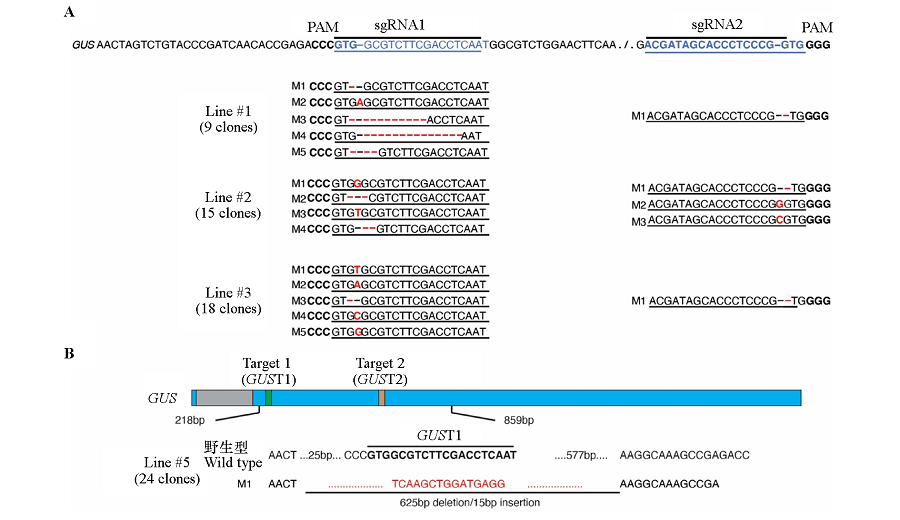
Fig. 4 The mutation types of #1,#2,#3(A)and #5(B)gene editing lines The blue bars represent the exons,the gray bar represents the intron,the green and orange bars represent two target sites,the numbers below the bar are position information,and the width of the bars are drawn in proportion to the sequence length. A:PAM sequence is shown in bold font;the target sites are shown in blue color;the mutations are represented in red color;B:The target sites are indicated by bold font,and the deletion and insertion are indicated by red color.
| [1] |
Bai S L, Tao R Y, Tang Y X, Yin L, Ma Y J, Ni J B, Yan X H, Yang Q S, Wu Z Y, Zeng Y L, Teng Y W. 2019. BBX16,a B-box protein,positively regulates light-induced anthocyanin accumulation by activating MYB10 in red pear. Plant Biotechnology Journal, 17(10):1985-1997.
doi: 10.1111/pbi.v17.10 URL |
| [2] |
Chen Y T, Mao W W, Liu T, Feng Q Q, Li L, Li B B. 2020. Genome editing as a versatile tool to improve horticultural crop qualities. Horticultural Plant Journal, 6(6):372-384.
doi: 10.1016/j.hpj.2020.11.004 URL |
| [3] | Cui Xia, Zhang Shuaibin. 2017. Gene editing technology and its application and prospect in horticultural crops. Acta Horticulturae Sinica, 44(9):1787-1795. (in Chinese) |
| 崔霞, 张率斌. 2017. 基因编辑技术及其在园艺作物中的应用和展望. 园艺学报, 44(9):1787-1795. | |
| [4] |
Du H Y, Zeng X R, Zhao M, Cui X P, Wang Q, Yang H, Cheng H, Yu D Y. 2016. Efficient targeted mutagenesis in soybean by TALENs and CRISPR/Cas9. Journal of Biotechnology, 217:90-97.
doi: 10.1016/j.jbiotec.2015.11.005 URL |
| [5] |
Graf R, Li X, Chu V T, Rajewsky K. 2019. sgRNA sequence motifs blocking efficient CRISPR/Cas9-mediated gene editing. Cell Reports, 26(5):1098-1103.
doi: 10.1016/j.celrep.2019.01.024 URL |
| [6] | Guo Ye, Wan Dongyan, Chai Zhuangzhuang, Wang Yuejin, Wen Yingqiang. 2019. Knock-out analysis of VviPDS1gene using CRISPR/Cas9 in grapevine . Acta Horticulturae Sinica, 46(4):623-634. (in Chinese) |
| 郭晔, 万东艳, 柴壮壮, 王跃进, 文颖强. 2019. 利用CRISPR/Cas9敲除葡萄VviPDS1基因的研究. 园艺学报, 46(4):623-634. | |
| [7] |
Jia H, Orbovic V, Jones J B, Wang N. 2016. Modification of the PthA4 effector binding elements in Type I CsLOB1 promoter using Cas9/sgRNA to produce transgenic Duncan grapefruit alleviating XccpthA4:dCsLOB1.3 infection. Plant Biotechnology Journal, 14(5):1291-1301.
doi: 10.1111/pbi.2016.14.issue-5 URL |
| [8] |
Jia H, Wang N. 2014. Targeted genome editing of sweet orange using Cas9/sgRNA. PLoS ONE, 9(4):e93806.
doi: 10.1371/journal.pone.0093806 URL |
| [9] |
Jia H, Zhang Y, Orbovic V, Xu J, White F F, Jones J B, Wang N. 2017. Genome editing of the disease susceptibility gene CsLOB1 in citrus confers resistance to citrus canker. Plant Biotechnology Journal, 15(7):817-823.
doi: 10.1111/pbi.12677 URL |
| [10] |
Li C, Unver T, Zhang B H. 2017. A high-efficiency CRISPR/Cas9 system for targeted mutagenesis in cotton(Gossypium hirsutumL.). Scientific Reports, 7:43902.
doi: 10.1038/srep43902 URL |
| [11] |
Li J, Li Y, Ma L G. 2019. CRISPR/Cas9-based genome editing and its applications for functional genomic analyses in plants. Small Methods, 3(3):1800473.
doi: 10.1002/smtd.v3.3 URL |
| [12] |
Li J F, Norville J E, Aach J, McCormack M, Zhang D D, Bush J, Church G M, Sheen J. 2013. Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nature Biotechnology, 31(8):688-691.
doi: 10.1038/nbt.2654 URL |
| [13] |
Ma C, Liu M, Li Q, Jun Si, Ren X, Song H. 2019. Efficient BoPDSgene editing in cabbage by the CRISPR/Cas9 system. Horticultural Plant Journal, 5(4):164-169.
doi: 10.1016/j.hpj.2019.04.001 URL |
| [14] | Ma X, Liu Y G. 2016. CRISPR/Cas9-based multiplex genome editing in monocot and dicot plants. Current Protocols in Molecular Biology,115:31.6.1-31. 6.21. |
| [15] | Malnoy M, Viola R, Jung M H, Koo O J, Kim S, Kim J S, Velasco R, Kanchiswamy C N. 2016. DNA-free genetically edited grapevine and apple protoplast using CRISPR/Cas9 ribonucleoproteins. Frontiers in Plant Science, 7:1904. |
| [16] |
Mao Y F, Zhang H, Xu N F, Zhang B T, Gou F, Zhu J K. 2013. Application of the CRISPR/Cas system for efficient genome engineering in plants. Molecular Plant, 6(6):2008-2011.
doi: 10.1093/mp/sst121 URL |
| [17] | Nakajima I, Ban Y, Azuma A, Onoue N, Moriguchi T, Yamamoto T, Toki S, Endo M. 2017. CRISPR/Cas9-mediated targeted mutagenesis in grape. PLoS ONE, 12(5):16. |
| [18] |
Nekrasov V, Staskawicz B, Weigel D, Jones J D G, Kamoun S. 2013. Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nature Biotechnology, 31(8):691-693.
doi: 10.1038/nbt.2655 URL |
| [19] |
Nishitani C, Hirai N, Komori S, Wada M, Okada K, Osakabe K, Yamamoto T, Osakabe Y. 2016. Efficient genome editing in apple using a CRISPR/Cas9 system. Scientific Reports, 6:31481.
doi: 10.1038/srep31481 URL |
| [20] |
Peng A, Chen S, Lei T, Xu L, He Y, Wu L, Yao L, Zou X. 2017. Engineering canker-resistant plants through CRISPR/Cas9-targeted editing of the susceptibility gene CsLOB1 promoter in citrus. Plant Biotechnology Journal, 15(12):1509-1519.
doi: 10.1111/pbi.2017.15.issue-12 URL |
| [21] |
Ren C, Liu X J, Zhang Z, Wang Y, Duan W, Li S H, Liang Z C. 2016. CRISPR/Cas9-mediated efficient targeted mutagenesis in Chardonnay(Vitis vinifera L.). Scientific Reports, 6:32289.
doi: 10.1038/srep32289 URL |
| [22] | Shan Qiwei, Gao Caixia. 2015. Recent advances in plant genome editing and its derivatives. Hereditas, 37(10):953-973. (in Chinese) |
| 单奇伟, 高彩霞. 2015. 植物基因组编辑及衍生技术最新研究进展. 遗传, 37(10):953-973. | |
| [23] |
Shan Q W, Wang Y P, Li J, Zhang Y, Chen K L, Liang Z, Zhang K, Liu J X, Xi J J, Qiu J L, Gao C X. 2013. Targeted genome modification of crop plants using a CRISPR-Cas system. Nature Biotechnology, 31(8):686-688.
doi: 10.1038/nbt.2650 URL |
| [24] |
Svitashev S, Young J K, Schwartz C, Gao H R, Falco S C, Cigan A M. 2015. Targeted mutagenesis,precise gene editing,and site-specific gene insertion in maize using Cas9 and guide RNA. Plant Physiology, 169(2):931-945.
doi: 10.1104/pp.15.00793 URL |
| [25] |
Tian S, Jiang L, Gao Q, Zhang J, Zong M, Zhang H, Ren Y, Guo S, Gong G, Liu F, Xu Y. 2017. Efficient CRISPR/Cas9-based gene knockout in watermelon. Plant Cell Reports, 36(3):399-406.
doi: 10.1007/s00299-016-2089-5 URL |
| [26] | Wang Fujun, Zhao Kaijun. 2018. Progress and challenge of crop genetic improvement via genome editing. Scientia Agricultura Sinica, 51(1):1-16. (in Chinese) |
| 王福军, 赵开军. 2018. 基因组编辑技术应用于作物遗传改良的进展与挑战. 中国农业科学, 51(1):1-16. | |
| [27] |
Wang X, Tu M, Wang D, Liu J, Li Y, Li Z, Wang Y, Wang X. 2018. CRISPR/Cas9-mediated efficient targeted mutagenesis in grape in the first generation. Plant Biotechnology Journal, 16(4):844-855.
doi: 10.1111/pbi.2018.16.issue-4 URL |
| [28] |
Wang Y P, Cheng X, Shan Q W, Zhang Y, Liu J X, Gao CX, Qiu J L. 2014. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nature Biotechnology, 32(9):947-951.
doi: 10.1038/nbt.2969 URL |
| [29] |
Wang Z, Wang S, Li D, Zhang Q, Li L, Zhong C, Liu Y, Huang H. 2018. Optimized paired-sgRNA/Cas9 cloning and expression cassette triggers high-efficiency multiplex genome editing in kiwifruit. Plant Biotechnology Journal, 16(8):1424-1433.
doi: 10.1111/pbi.2018.16.issue-8 URL |
| [30] |
Wu J, Wang Z, Shi Z, Zhang S, Ming R, Zhu S, Khan M, Tao S, Korban S, Wang H, Chen N, Nishio T, Xu X, Cong L, Qi K, Huang X, Wang Y, Zhao X, Wu J, Deng C, Gou C, Zhou W, Yin H, Qin G, Sha Y, Tao Y, Chen H, Yang Y, Song Y, Zhan D, Wang J, Li L, Dai M, Gu C, Wang Y, Shi D, Wang X, Zhang H, Zeng L, Zheng D, Wang C, Chen M, Wang G, Xie L, Sovero V, Sha S, Huang W, Zhang S, Zhang M, Sun J, Xu L, Li Y, Liu X, Li Q, Shen J, Wang J, Paull R, Bennetzen J, Zhang S. 2013. The genome of the pear(Pyrus bretschneideriRehd.). Genome Research, 23:396-408.
doi: 10.1101/gr.144311.112 URL |
| [31] | Yang Lushan, Guo Ye, Hu Yang, Wen Yingqiang. 2020. CRISPR/Cas9-mediated mutagenesis of VviEDR2 results in enhanced resistance to powdery mildew in grapevine (Vitis vinifera) . Acta Horticulturae Sinica, 47(4):623-634. (in Chinese) |
| 杨禄山, 郭晔, 胡洋, 文颖强. 2020. 利用CRISPR/Cas9系统敲除葡萄中VviEDR2提高对白粉病的抗性. 园艺学报, 47(4):623-634. | |
| [32] |
Yang Q, Yang B, Li J, Wang Y, Tao R, Yang F, Wu X, Yan X, Ahmad M, Shen J, Bai S, Teng Y. 2020. ABA-responsive ABRE-BINDING FACTOR3 activates DAM3 expression to promote bud dormancy in Asian pear. Plant Cell and Environment, 43(6):1360-1375.
doi: 10.1111/pce.v43.6 URL |
| [33] | Zheng Aihong, Zhang Fen, Jiang Min, Yuan Qiao, Jiang Leiyu, Chen Qing, Tang Haoru, Sun Bo. 2019. Targeted editing of BoaZDSby CRISPR/Cas9 technology in Chinese kale . Acta Horticulturae Sinica, 46(1):57-64. (in Chinese) |
| 郑爱红, 张芬, 江敏, 袁巧, 江雷雨, 陈清, 汤浩茹, 孙勃. 2019. 利用CRISPR/Cas9技术靶向编辑芥蓝BoaZDS. 园艺学报, 46(1):57-64. | |
| [34] | Zou Xiuping, Fan Di, Peng Aihong, He Yongrui, Xu Lanzhen, Lei Tiangang, Yao Lixiao, Li Qiang, Luo Keming, Chen Shanchun. 2019 CRISPR/Cas9-mediated editing of multiple sites in the citrus CsLOB1 promoter . Acta Horticulturae Sinica, 46(2):337-344. (in Chinese) |
| 邹修平, 范迪, 彭爱红, 何永睿, 许兰珍, 雷天刚, 姚利晓, 李强, 罗克明, 陈善春. 2019. CRISPR/Cas9介导柑橘CsLOB1基因启动子的多位点编辑. 园艺学报, 46(2):337-344. |
| [1] | QIANG Wenyan, MENG Qingran, ZHANG Zhiguo, GAO Wenjie. Analysis of Petal Volatile Components Among Different Hemerocallis Cultivars Based on HS-SPME-GC-MS [J]. Acta Horticulturae Sinica, 2023, 50(1): 116-130. |
| [2] | SONG Jiankun, YANG Yingjie, LI Dingli, MA Chunhui, WANG Caihong, and WANG Ran. A New Pear Cultivar‘Luxiu’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 3-4. |
| [3] | DONG Xingguang, CAO Yufen, ZHANG Ying, TIAN Luming, HUO Hongliang, QI Dan, XU Jiayu, LIU Chao, and WANG Lidong. A New Cold-resistant Crispy Pear Cultivar‘Yucuixiang’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 5-6. |
| [4] | OU Chunqing, JIANG Shuling, WANG Fei, MA Li, ZHANG Yanjie, and LIU Zhenjie. A New Early-ripening Pear Cultivar‘Xingli Mishui’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 7-8. |
| [5] | ZHANG Yanjie, WANG Fei, OU Chunqing, MA Li, JIANG Shuling, and LIU Zhenjie. A New Pear Cultivar‘Zhongli Yucui 3’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 9-10. |
| [6] | WANG Suke, LI Xiugen, YANG Jian, WANG Long, SU Yanli, ZHANG Xiangzhan, and XUE Huabai. A New Red Pear Cultivar‘Danxiahong’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 13-14. |
| [7] | YAN Lei, FAN Lijuan, MEN Wanjie, and LIU Zhiyang, . A New Hemerocallis Cultivar‘Xuan Cai Jing Ling’ [J]. Acta Horticulturae Sinica, 2022, 49(S2): 199-200. |
| [8] | WANG Fei, OU Chunqing, ZHANG Yanjie, MA Li, and JIANG Shuling. A New Late Ripening Pear Cultivar‘Huaqiu’with Long Storage Period [J]. Acta Horticulturae Sinica, 2022, 49(S1): 9-10. |
| [9] | SONG Jiankun, LI Dingli, YANG Yingjie, MA Chunhui, WANG Caihong, and WANG Ran. A New Pear Cultivar‘Qindaohong’ [J]. Acta Horticulturae Sinica, 2022, 49(S1): 11-12. |
| [10] | GUO Weizhen, ZHAO Jingxian, LI Ying. A New Mid-early Ripening Pear Cultivar‘Meiyu’ [J]. Acta Horticulturae Sinica, 2022, 49(9): 2051-2052. |
| [11] | LIU Jinming, GUO Caihua, YUAN Xing, KANG Chao, QUAN Shaowen, NIU Jianxin. Genome-wide Identification of Dof Family Genes and Expression Analysis Sepal Persistent and Abscission in Pear [J]. Acta Horticulturae Sinica, 2022, 49(8): 1637-1649. |
| [12] | TAO Xin, ZHU Rongxiang, GONG Xin, WU Lei, ZHANG Shaoling, ZHAO Jianrong, ZHANG Huping. Fructokinase Gene PpyFRK5 Plays an Important Role in Sucrose Accumulation of Pear Fruit [J]. Acta Horticulturae Sinica, 2022, 49(7): 1429-1440. |
| [13] | ZUO Xin, LI Mingming, LI Xinrong, MIAO Chunyan, LI Yanfang, YANG Xu, ZHANG Zhongyi, WANG Fengqing. CRISPR/Cas9 Technology for RcPDS1 Gene Editing in Rehmannia chingii [J]. Acta Horticulturae Sinica, 2022, 49(7): 1532-1544. |
| [14] | LIANG Qin, ZHANG Yanhui, KANG Kaiquan, LIU Jinhang, LI Liang, FENG Yu, WANG Chao, YANG Chao, LI Yongyu. Molecular Evolution of MiR168 Family and Their Expression Profiling During Dormancy of Pyrus pyrifolia [J]. Acta Horticulturae Sinica, 2022, 49(5): 958-972. |
| [15] | XIANG Miaolian, WU Fan, LI Shucheng, MA Qiaoli, WANG Yinbao, XIAO Liuhua, CHEN Jinyin, CHEN Ming. Exogenous Melatonin Regulates Reactive Oxygen Metabolism to Induce Resistance of Postharvest Pear Fruit to Black Spot [J]. Acta Horticulturae Sinica, 2022, 49(5): 1102-1110. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
Copyright © 2012 Acta Horticulturae Sinica 京ICP备10030308号-2 国际联网备案号 11010802023439
Tel: 010-82109523 E-Mail: yuanyixuebao@126.com
Support by: Beijing Magtech Co.Ltd